Latent semantic indexing (LSI) was first introduced for the analysis of scATAC-seq data by Cusanovich et al. 2018[1]. LSI combines steps of frequency-inverse document frequency (TF-IDF) normalization followed by singular value decomposition (SVD). Partek Flow wrapped Signac's TF-IDF normalization for single cell ATAC-seq dataset. It is a two-step normalization procedure that both normalizes across cells to correct for differences in cellular sequencing depth, and across peaks to give higher values to more rare peaks[2].
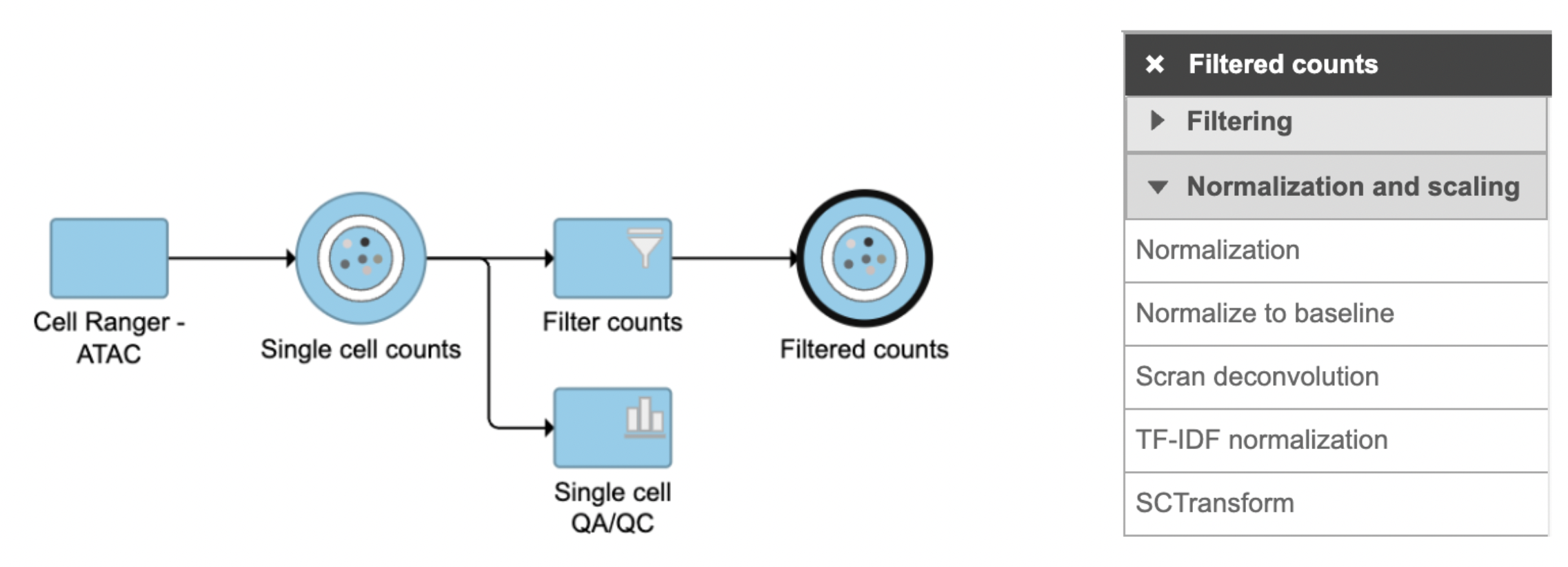
TF-IDF normalization in Flow can be invoked in Normalization and scaling section by clicking any single cell counts data node (Figure 1).
Figure 1. TF-IDF normalization task in Normalization and scaling section in Flow.
To run TF-IDF normalization,
- Click a single cell counts data node
- Click the Normalization and scaling section in the toolbox
- Click TF-IDF normalization
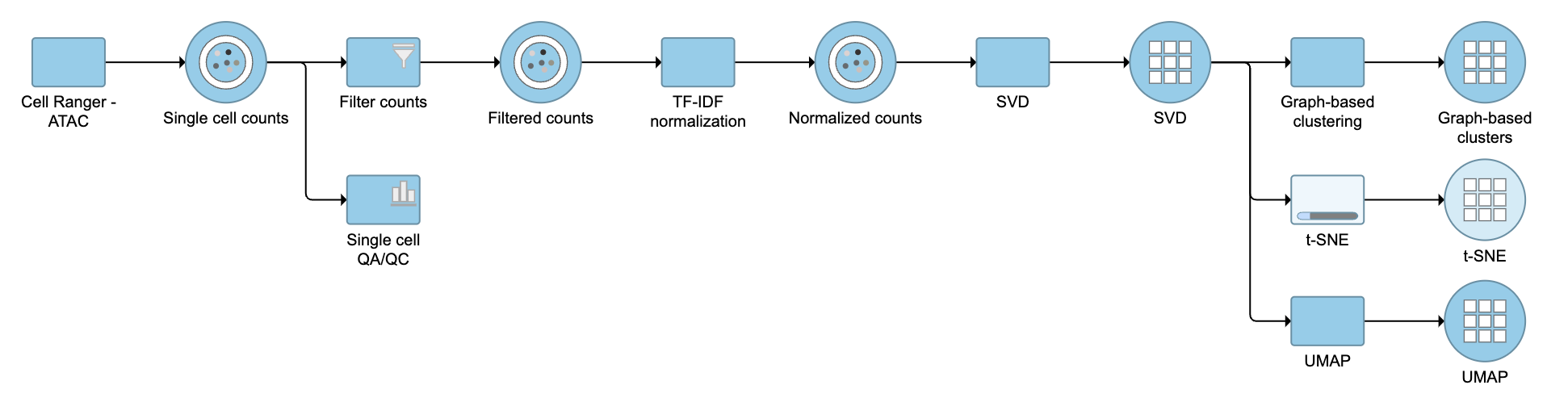
The output of TF-IDF normalization is a new data node that has been normalized by log(TF x IDF). We can then use this new normalized matrix for downstream analysis and visualization (Figure 2).
Figure 2. Example workflows to demonstrate downstream analysis and visualization of TF-IDF normalization output.
References
- Cusanovich, D., Reddington, J., Garfield, D. et al. The cis-regulatory dynamics of embryonic development at single-cell resolution. Nature 555, 538–542 (2018). https://doi.org/10.1038/nature25981
- https://satijalab.org/signac/index.html
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.