Join us for an event September 26!
How to Streamline RNA-Seq analysis and increase productivity—point, click, and done
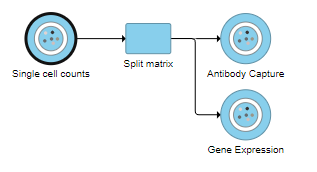
Split matrix can be invoked on any counts data node with more than one feature type. For example, a CITE-Seq experiment would have Gene Expression counts and Antibody Capture counts in the single cell counts data node. Datasets generated by 10X Genomics' Feature Barcoding experiments also utilize this task to split different feature measurements for downstream analysis.
There are no parameters to configure, to run:
- Click the counts data node you want to split
- Click the Pre-analysis tools section of the toolbox
- Click Split matrix
The Split matrix task will run and generate output data nodes for each of the feature types. For example, if there are Antibody Capture and Gene Expression feature types in the input, Split matrix will generate two data nodes (Figure 1). Every sample is included in both matrices.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
29 | rates |