Seurat v3[1] introduced new methods for the integration of multiple single-cell datasets, no matter whether they were collected from different individuals, experimental conditions, technologies, etc. Seurat 3 integration method aims to use a subset of the data as reference for the integrate analysis. The method integrates all other data with the reference subset. The subset can be one sample or a subgroup of samples defined by the factor attribute.
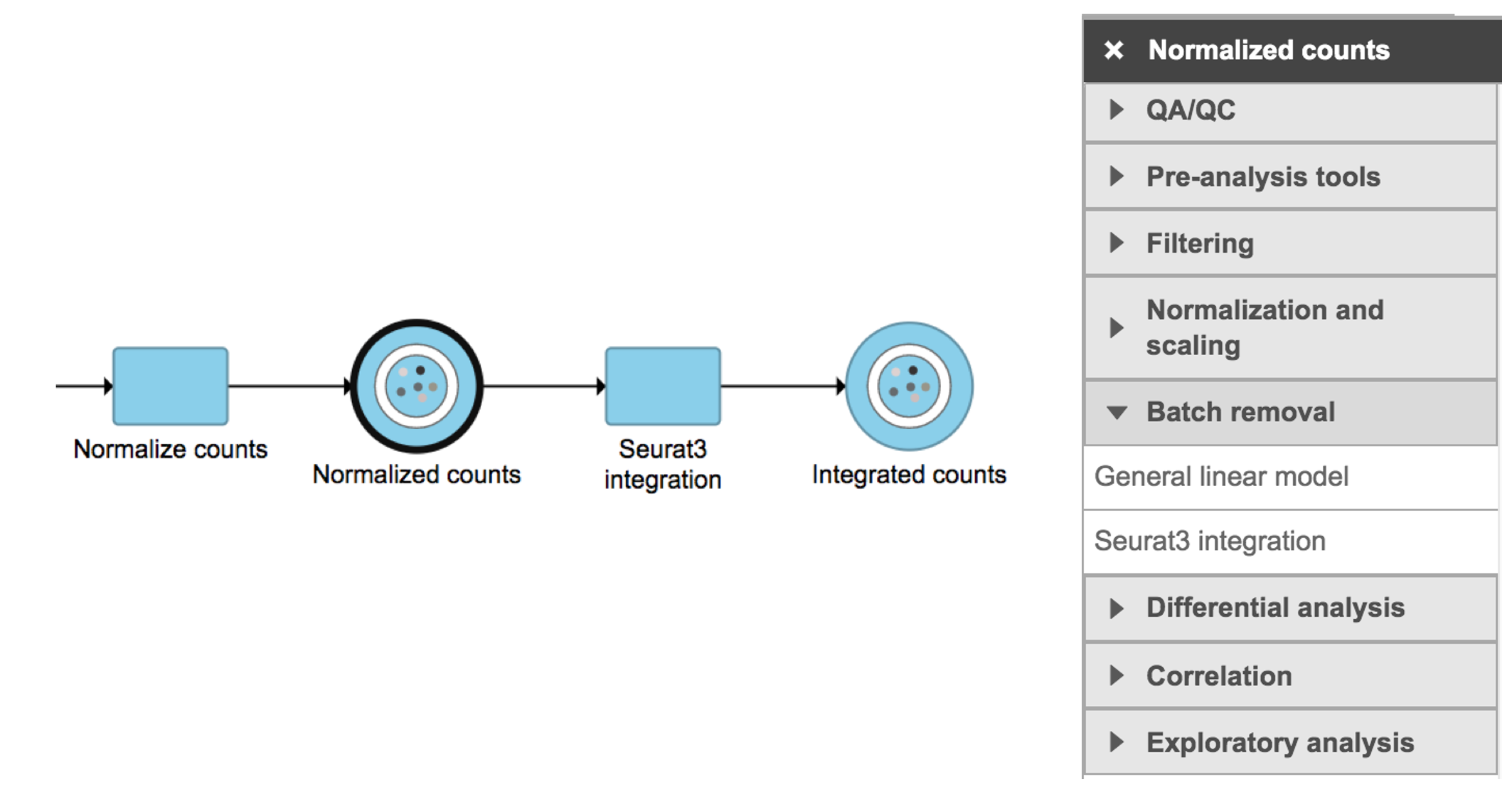
Seurat3 integration in Flow can be invoked in Batch removal section if a Normalized counts data node is selected (Figure 1).
Figure 1. Seurat3 integration task in Batch removal section in Flow.
To run Seurat3 integration,
- Click a Normalized counts data node
- Click the Batch removal section in the toolbox
- Click Seurat3 Integration
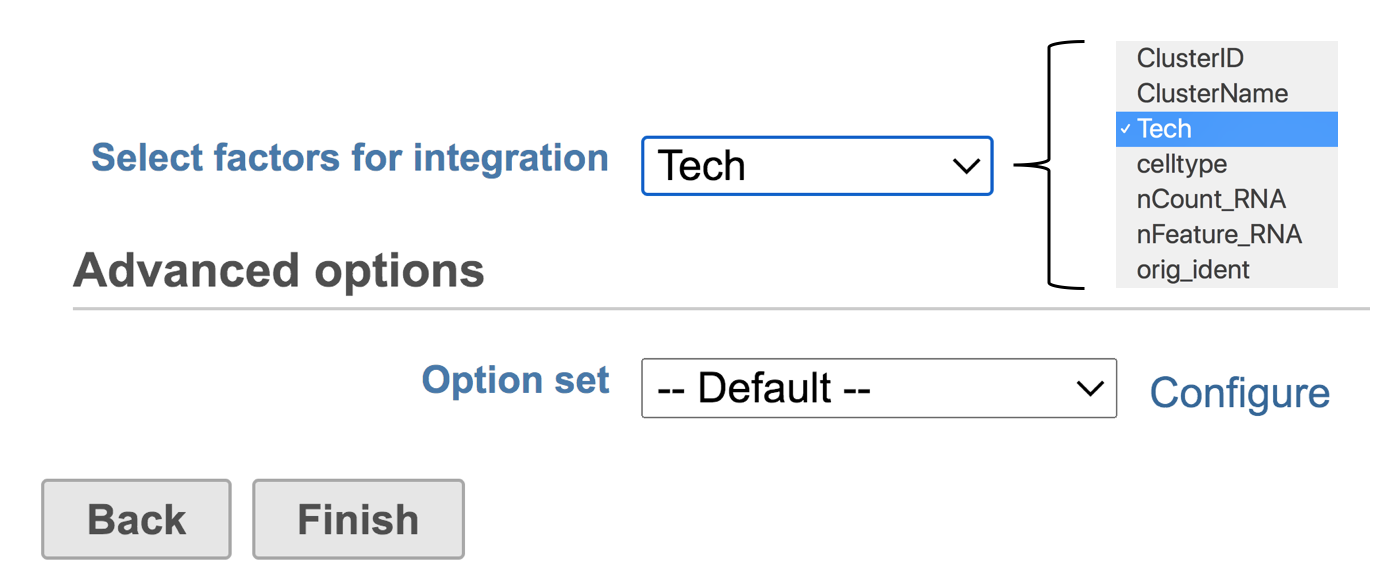
You will be promoted to pick some attribute(s) for analysis. The first Seurat3 integration dialog is a drop-down list that includes the factors for data integration. To set up the model, you need to choose which attribute should be considered. For example, in the case of one dataset that has different cell types from multiple technologies(Tech), different technology may have divergent impacts on different cell types. Hence, the attribute Tech should be considered to be the batch factor. The attribute celltype represents different cell types in this dataset (Figure 2).
To integrate data with default settings,
- Select Tech from the dropdown list
- Click Finish
Figure 2. Interface of Seurat3 integration in Flow. Example attributes are indicated in the drop-down list.
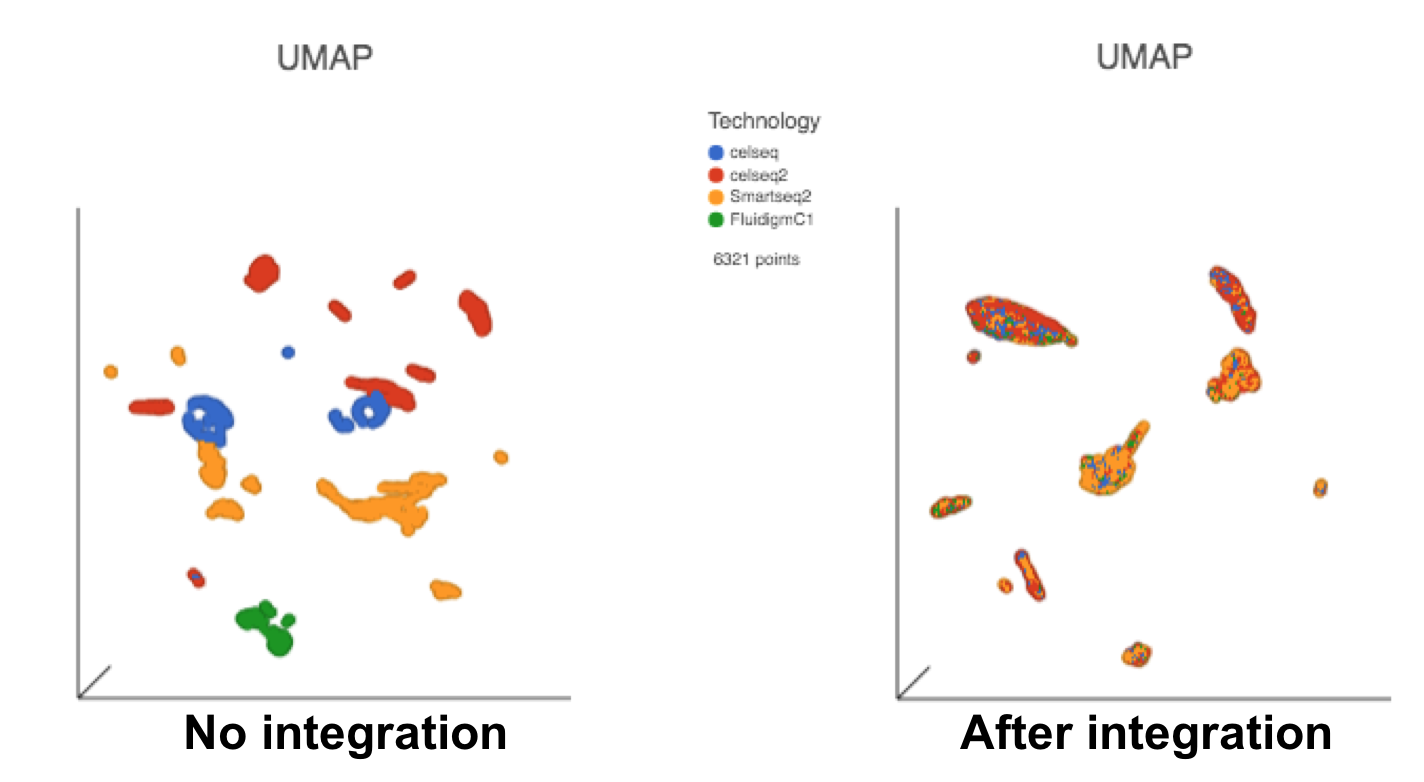
The output of Seurat3 integration is a new data node - Integrated counts (Figure 1). We can then use this new integrated matrix for downstream analysis and visualization (Figure 3).
Figure 3. UMAP displays the cells colored by Tech attribute before(left) and after(right) Seurat3 integration. Both images in the figure share the same legend.
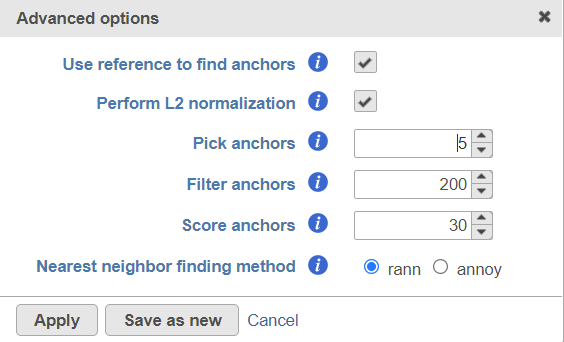
Users can click Configure to change the default settings In Advanced options (Figure 4).
Figure 4. Advanced configure options for Seurat3 integration in Flow.
Use reference to find anchors: when this box is checked, the first group of the selected attribute is used as reference to find anchors. To use a different group as reference, change the order of subgroups of the attribute in the attribute management page on Data tab. When the box is unchecked, anchors will be identified by comparing all pairs of subgroups, this option is very computationally intensive.
Perform L2 normalization: Perform L2 normalization on the CCA cell embeddings after dimensional reduction.
Pick anchors: How many neighbors (k) to use when picking anchors.
Filter anchors: How many neighbors (k) to use when filtering anchors.
Score anchors: How many neighbors (k) to use when scoring anchors.
Nearest neighbour finding methods: Method for nearest neighbor finding. Options include: rann, annoy.
References
- Stuart T, Butler A, Hoffman P, et al. Comprehensive integration of single-cell data. Cell, 2019. DOI:https://doi.org/10.1016/j.cell.2019.05.031
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.