The Kruskal-Wallis and Dunn's tests (Non-parametric ANOVA) task is used to identify deferentially expressed genes among two or more groups. Note that such rank-based tests are generally advised for use with larger sample sizes.
Running the task
To invoke the Kruskal-Wallis test, select any count-based data nodes, these include:
- Gene counts
- Transcript counts
- Normalized counts
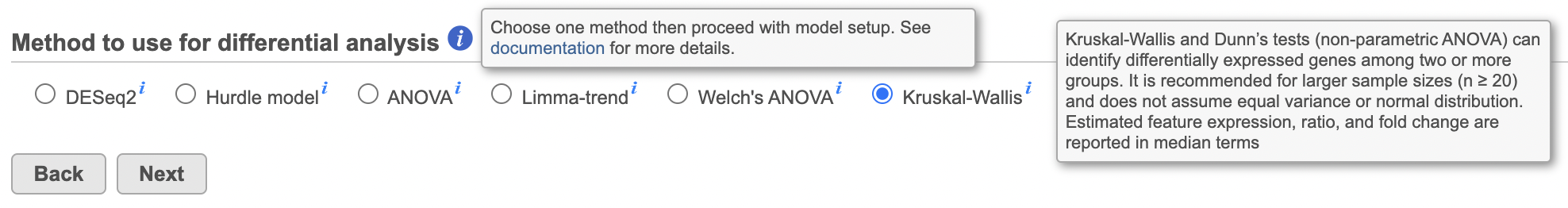
Select Statistics > Differential analysis in the context-sensitive menu, then select Kruskal-Wallis (Figure 1).
Select a specific factor for analysis and click the Next button (Figure 2). Note that this task can only take into account one factor at a time.
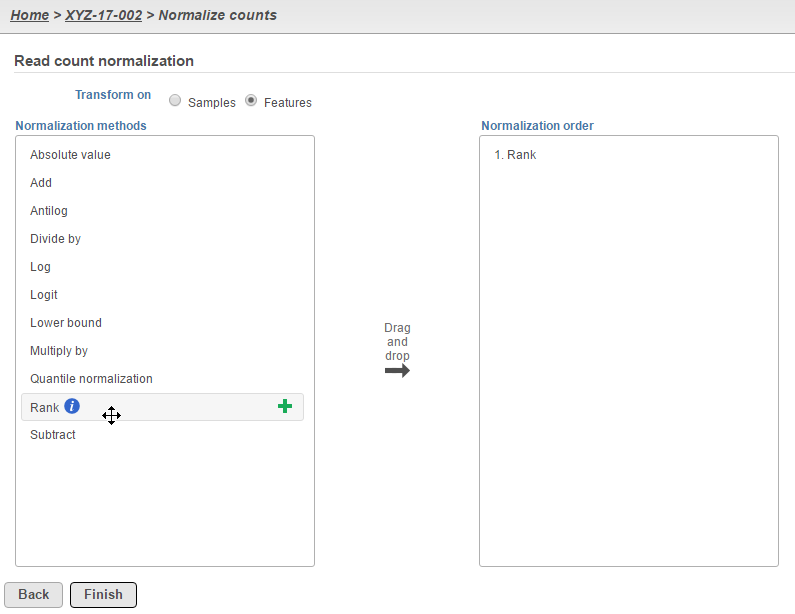
For more complicated experimental designs, go back to the original count data that will be used as input and perform Rank normalization at the Features level (Figure 3). The resulting Normalized counts data node can then be analyzed using the Detect differential expression (ANOVA) task, which can take into account multiple factors as well as interactions.
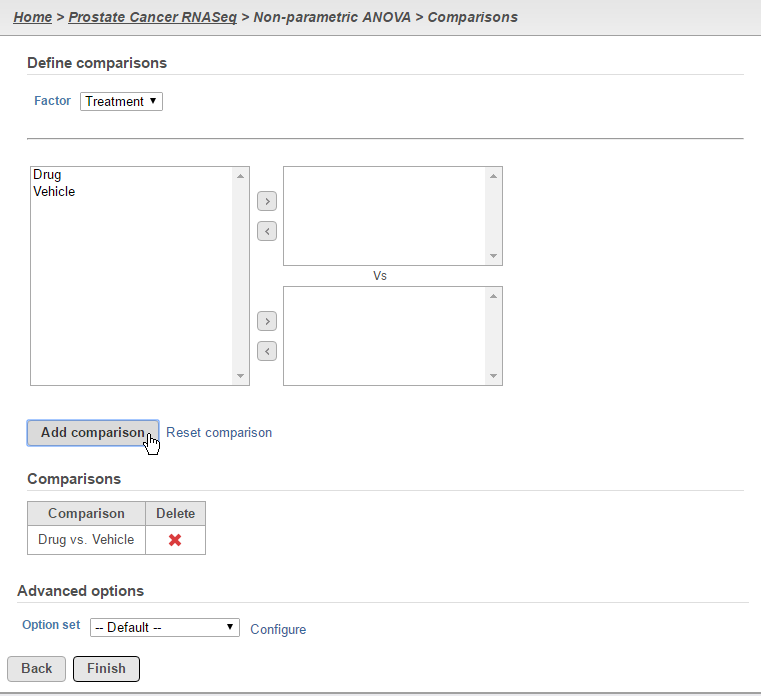
Define the desired comparisons between groups and click the Finish button (Figure 4). Note that comparisons can only be added between single group (i.e. one group per box).
Report
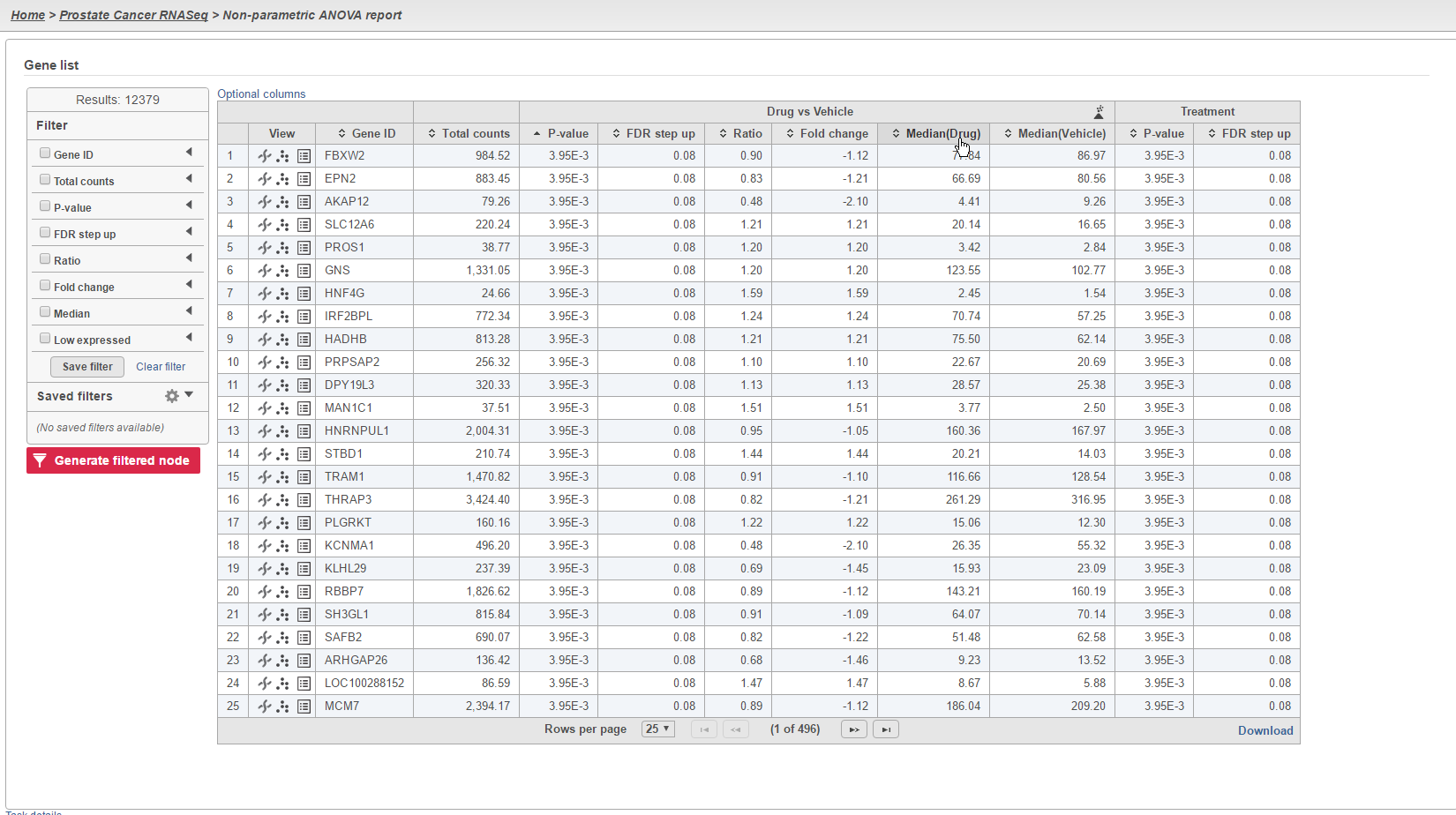
The results of the analysis will appear similar to other differential expression analysis results. However, the column to indicate mean expression levels for each group will display the median instead (Figure 5).
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
41 | rates |