What is K-means clustering?
K-means clustering is a method for identifying groups of similar observations, i.e. cells or samples. K-means clustering aims to group observations into a pre-determined number of clusters (k) so that each observation belongs to the cluster with the nearest mean. An important aspect of K-means clustering is that it expects clusters to be of similar size (equal variance) and shape (distribution of variance is spherical). The Compare Clusters task can also be used to help determine the optimal number of K-means clusters.
Running K-means clustering
We recommend normalizing your data prior to running K-means clustering, but the task will run on any counts data node.
- Click the counts data node
- Click the Exploratory analysis section of the toolbox
- Click K-means clustering
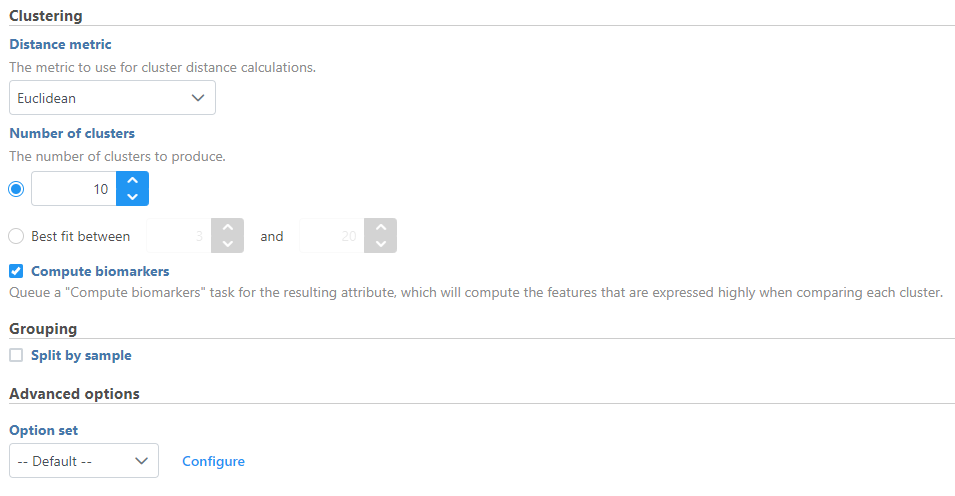
- Configure the parameters
- Click Finish to run (Figure 1)
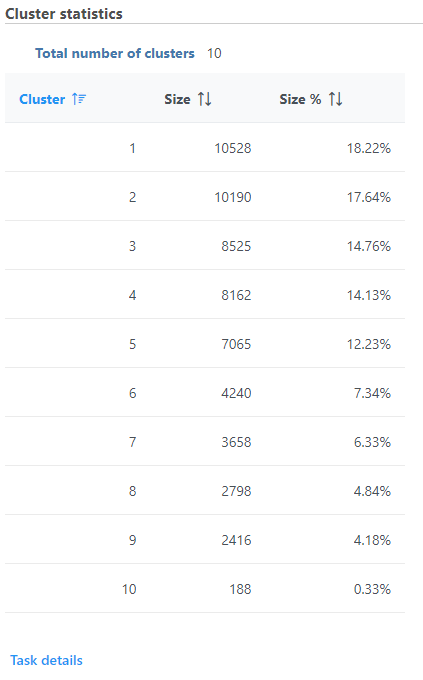
Cluster statistics
The total number of clusters is listed along with the number and percentage of cells in each cluster.
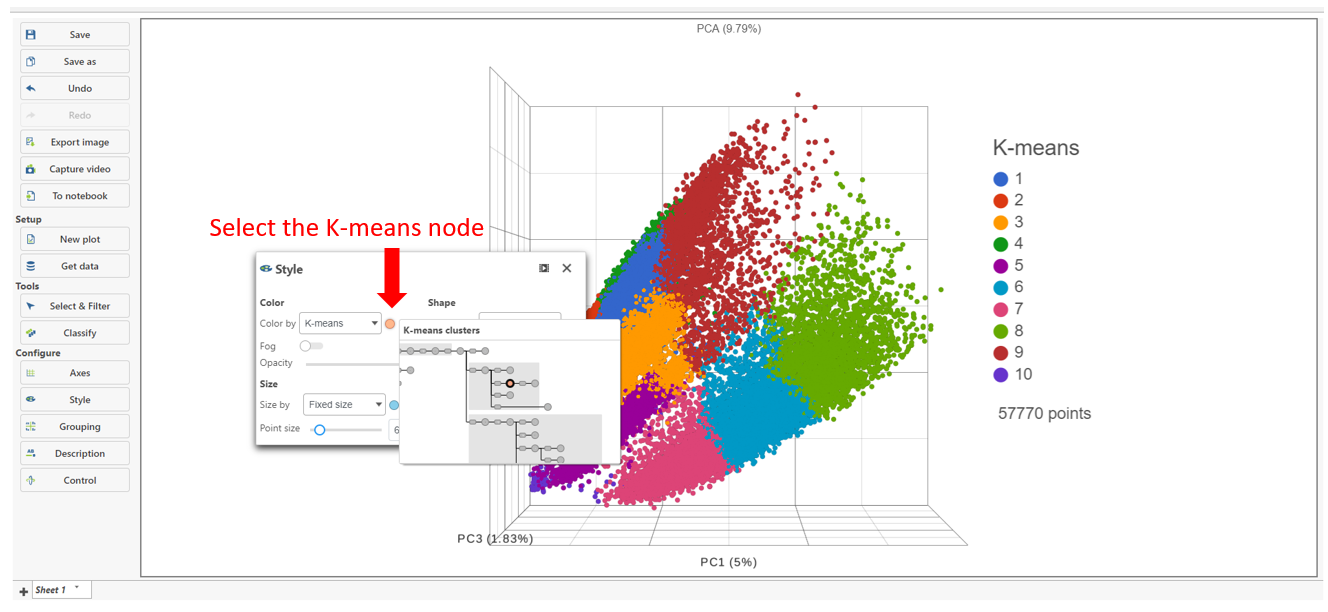
The K-means Clustering result data node includes the input values for each gene and adds cluster assignment as a new attribute, K-means, for each observation. If the K-means clusters data node is visualized by Scatter plot, PCA, t-SNE, or UMAP, the plot can be colored by the K-means attribute (Figure 3).
Basic K-means clustering parameters
Distance metric
Choose which distance metric to use for cluster distance calculations. Options include Euclidean, Absolute Value, Euclidean Squared, Kendall Correlation, Max Value, Min Value, Pearson Correlation, Rank Correlation, Average Euclidean, Shape, Cosine, Canberra, Bray Curtis, Tanimoto, Pearson Correlation Absolute, Rank Correlation Absolute, and Kendall Correlation Absolute. The default is Euclidean.
Number of clusters
Choose between specifying a set number of clusters or a range to test for the best fit number of clusters. The best fit is determined by the number of clusters with the lowest Davies–Bouldin index. The default is set to 10 for a fixed number of clusters. The initial values for the range option are 3 to 20 clusters.
Compute biomarkers
Choose whether to run the ANOVA test comparing each cluster to all other observations to identify features that have higher values in that cluster. Default is Enabled.
Split cells by sample
This option is present in single cell data. If enabled, K-means clustering will be run separately for each sample. If disabled, K-means clustering will be run on all cells from the input data. Default is set by the Split single cell by sample option in the user preference page.
Advanced K-means clustering parameters
Random cluster initialization
If enabled, the initial cluster centroids will be selected randomly from among the data points. If disabled, the initial cluster centroids will be selected to optimize distance between clusters. Default is Disabled.
Random seed
This sets the random seed used if Random cluster initialization is enabled. Use the same random seed to reproduce results.
Batch centroid computations
If enabled, all cluster centroids will be recomputed at the end of each iteration. If disabled, each cluster centroid will be recomputed as the members of the cluster change. Default is Enabled.
Max iterations
The maximum number of iterations to perform before setting on a set of clusters. Default is 1000.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
22 | rates |