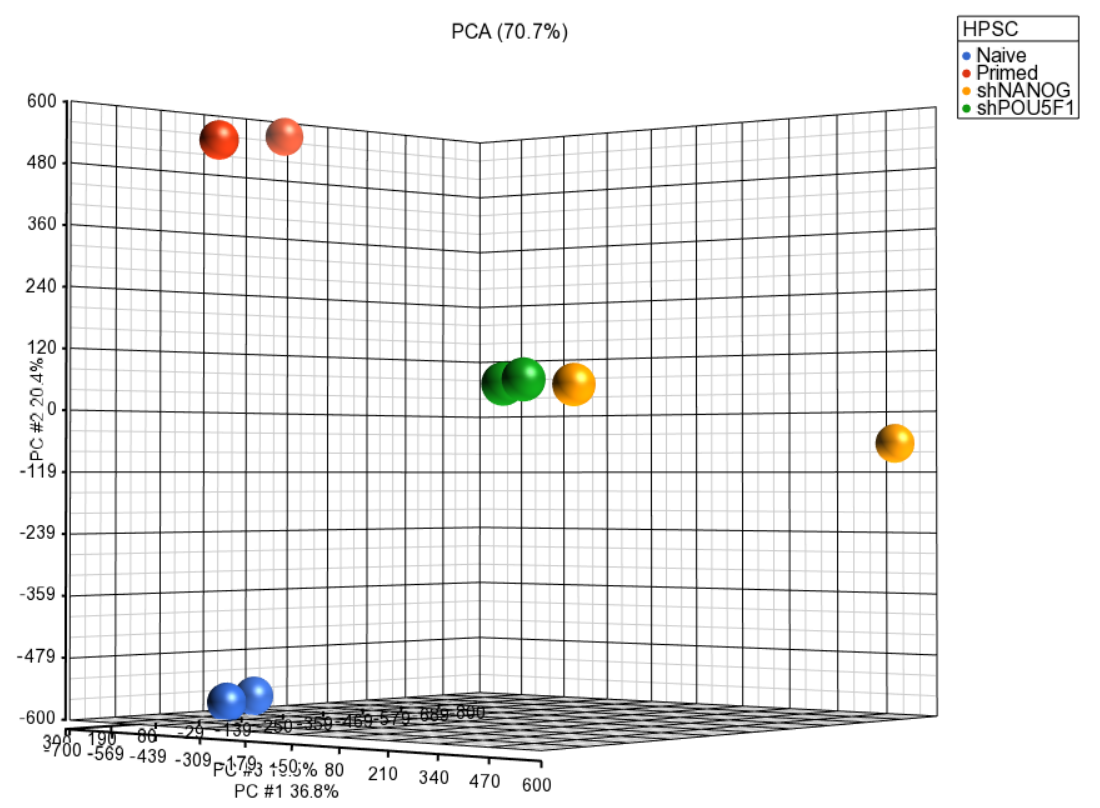
Principal component analysis (PCA) can be invoked on the methylation data to reveal clustering of the samples, but also as a quality control procedure (detection of outliers could point to possible low quality or mislabeled samples). To obtain the PCA plot, switch to the Scatter Plot tab, push Recompute ( ) and from the Color by drop down list select HPSC. Use the Rotate Mode ( )to explore the plot from different angles, as shown in Figure 1. Each dot of the plot is a single sample and represents the average methylation status across all CpG loci. The result is shown in the demonstrating clear separation of naive and primed HPSC from the cells .
Section Heading
Section headings should use level 2 heading, while the content of the section should use paragraph (which is the default). You can choose the style in the first dropdown in toolbar.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |