Single cell RNA-seq gene expression counts are zero inflated due to inefficient mRNA capture. This normalization task is based on MAGIC[1]–MArkov Affinity-based Graph Imputation of Cells), to recover gene expression lost due to drop-out. The limitation on using this method is up to 50K cells in the input data node.
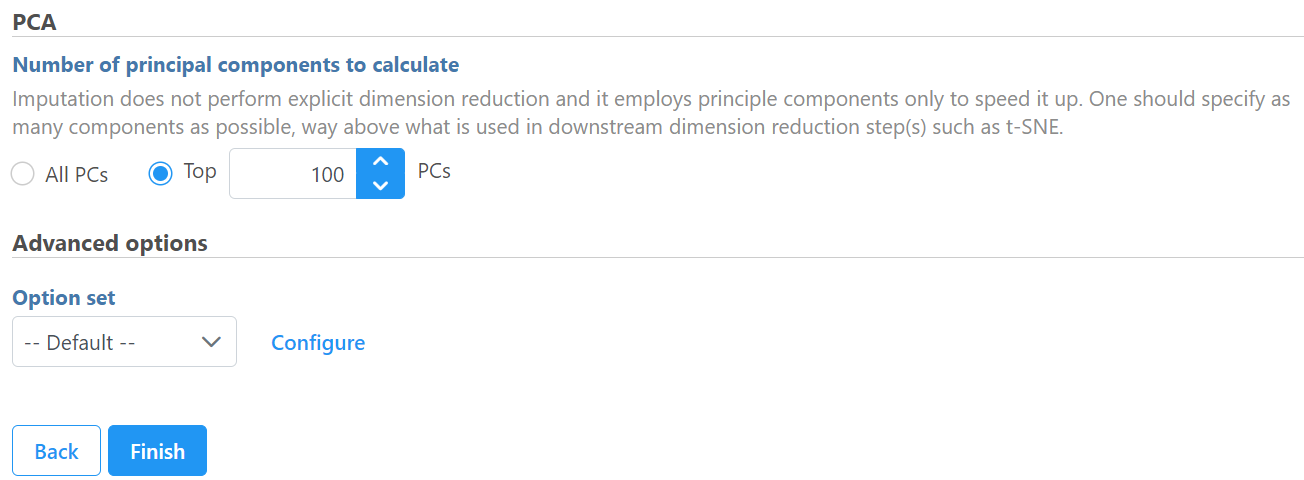
To invoke this task, click on a normalized data node which has less than 50K cells, it will first compute PCA to use the number of PCs specified to impute.
|
Click Finish to run the task, it will output low expression imputed matrix in the output report node.
References