With the Partek Flow REST API, you can create custom solutions to query or drive your server. Below are some common use cases for the REST API:
A complete reference for the API can be found on the REST API Command List or by visiting [server]/api/v1/servlets
The referenced Python library can be downloaded here.
An access token can be generated from the System information section of the settings page.

Alternatively, GetToken.py will generate a token:
python GetToken.py --server localhost:8080 --user admin |
you will be prompted to enter your password.
This token can be specified as the token parameter.
curl --form token=cUOWY0VvkSFagr... http://localhost:8080/flow/api/v1/users/list |
Flow organizes data by projects and they can be created and managed by the REST API.
To create a project:
curl -X POST --form token=$FLOW_TOKEN --form project="My Project" http://localhost:8080/flow/api/v1/projects |
The server will respond with JSON data describing the new project:
{"name":"My Project","id":"0","description":"","owner":"0","userRoles":{"0":"Project owner"},"outputFolders":{"0":"/home/flow/FlowData/Project_My Project"},"diskUsage":"0 GB","lastModifiedTimeStamp":1506013662476,"lastModifiedDate":"12:00 PM","data":[]} |
The new project will appear on the Flow homepage:
UploadSamples.py is a python script that can create samples within a project by uploading files:
python UploadSamples.py --verbose --token $FLOW_TOKEN --server http://localhost:8080 --project "My Project" \ --files ~/MoreData/REST/sample1.fastq.gz ~/MoreData/REST/sample2.fastq.gz ~/MoreData/REST/sample3.fastq.gz ~/MoreData/REST/sample4.fastq.gz |
This operation will generate a data node on the Analyses tab for the imported samples:
We can associate attributes with samples for use in visualizations and statistical analysis:
python AddAttribute.py -v --server http://localhost:8080 --token $FLOW_TOKEN --project_name "My Project" --sample_name sample1 --attribute Type --value Case python AddAttribute.py -v --server http://localhost:8080 --token $FLOW_TOKEN --project_name "My Project" --sample_name sample2 --attribute Type --value Case python AddAttribute.py -v --server http://localhost:8080 --token $FLOW_TOKEN --project_name "My Project" --sample_name sample3 --attribute Type --value Control python AddAttribute.py -v --server http://localhost:8080 --token $FLOW_TOKEN --project_name "My Project" --sample_name sample4 --attribute Type --value Control |
The sample attributes can be viewed and managed on the data tab:
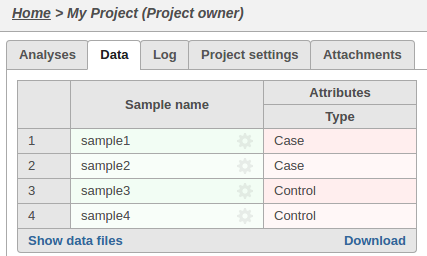
A pipeline is a series of tasks used to process and analyze genomic data. You can read more about pipelines here
To run a pipeline, first we need to know its name.
We can get the name of a pipeline from the GUI or from the API:
wget -q -O - http://localhost:8080/flow/api/v1/pipelines/list$AUTHDETAILS | python -m json.tool | gvim - |
Many pipelines also require that library files are specified.
You can get the list of required inputs for the pipeline from the API:
http://localhost:8080/flow/api/v1/pipelines/inputs?project_id=0&pipeline=AlignAndQuantify
This particular pipeline requires a bowtie index and an annotation model:

The request to launch the pipeline needs to specify one resource ID for each input.
These IDs can be found using the API:
Get the IDs for the library files that match the required inputs
wget -q -O - "http://localhost:8080/flow/api/v1/library_files/list${AUTHDETAILS}&assembly=hg19" | python -m json.tool | gvim - |
[
{
"annotationModel": "",
"assembly": "hg19",
"description": "Reference sequence",
"fileType": "Genome sequence",
"id": 100
},
{
"annotationModel": "",
"assembly": "hg19",
"description": "Cytoband",
"fileType": "cytoBand.txt",
"id": 101
},
{
"annotationModel": "",
"assembly": "hg19",
"description": "Bowtie index",
"fileType": "Bowtie Index",
"id": 102
},
{
"annotationModel": "hg19_refseq_15_05_07_v2",
"assembly": "hg19",
"description": "Annotation file: hg19_refseq_15_05_07_v2",
"fileType": "Annotation model",
"id": 103
}
] |
The pipeline can be launched in any project using RunPython.py
python RunPipeline.py -v --server http://localhost:8080 --token $FLOW_TOKEN --project_id 0 --pipeline AlignAndQuantify --inputs 102,103 |
This action will cause two tasks to start running:
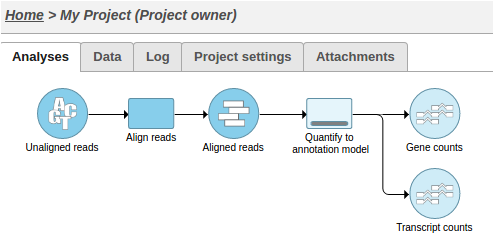
Alternatively, UploadSamples.py can create the project, upload the samples and launch the pipeline in one step:
python UploadSamples.py -v --server http://localhost:8080 --token $FLOW_TOKEN --files ~/sampleA.fastq.gz ~/sampleB.fastq.gz --project NewProject --pipeline AlignAndQuantify --inputs 102,103 |
To add a collaborator to a project:
curl -X PUT "http://localhost:8080/flow/api/v1/projects?project=ProjectName&collaborator=user1&role=Collaborator&token=$FLOW_TOKEN" |
curl --form token=$TO_TOKEN --form url=http://from:8080/flow/api/v1/feature_lists/export?token=$FROM_TOKEN http://to:8080/flow/api/v1/feature_lists/import |
#!/bin/bash
inotifywait -m $PATH_TO_MONITOR -e create -e moved_to |
while read path action file; do
if [[ $file == *.fastq.gz ]]; then
echo "Uploading $file"
python UploadSamples.py -v --server $SERVER --token $FLOW_TOKEN --files $path/$file --project "$PROJECT"
fi
done |
#!/bin/bash
while true; do
result=`python QueueStatistics.py --server $SERVER --token $TOKEN --max_waiting $MAX_WAITING`
if [ $? -eq 1 ]; then
/usr/bin/notify-send $result
exit 1
fi
sleep $INTERVAL
done |