Page History
...
When Other is selected, a specific value of the effective genome size needs to be specified in bps as unit (Figure 2).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
When sample attribute is not specified, for instance there are only two sample -- ChIP and mock as sample name, the peak detection pairs needs to be manually defined (Figure 1).
...
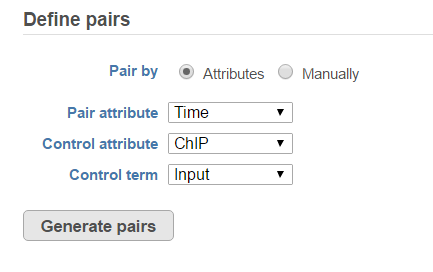
When select MACS2 task, the default option is to use sample attribute to add multiple pairs at one button click (Figure 4)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
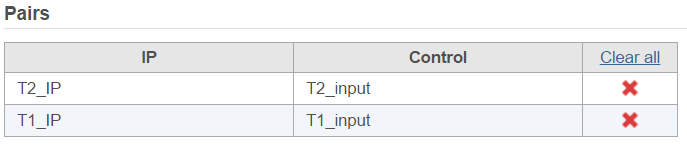
There are IP-Input pair in each time point, so the pair attribute is Time; Control attribute is the attribute contains IP and input group, which is ChIP, the control term is labeled as Input in the example, when click Generate pairs, the two pairs will be automatically added to the Pairs table at once (Figure 5).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
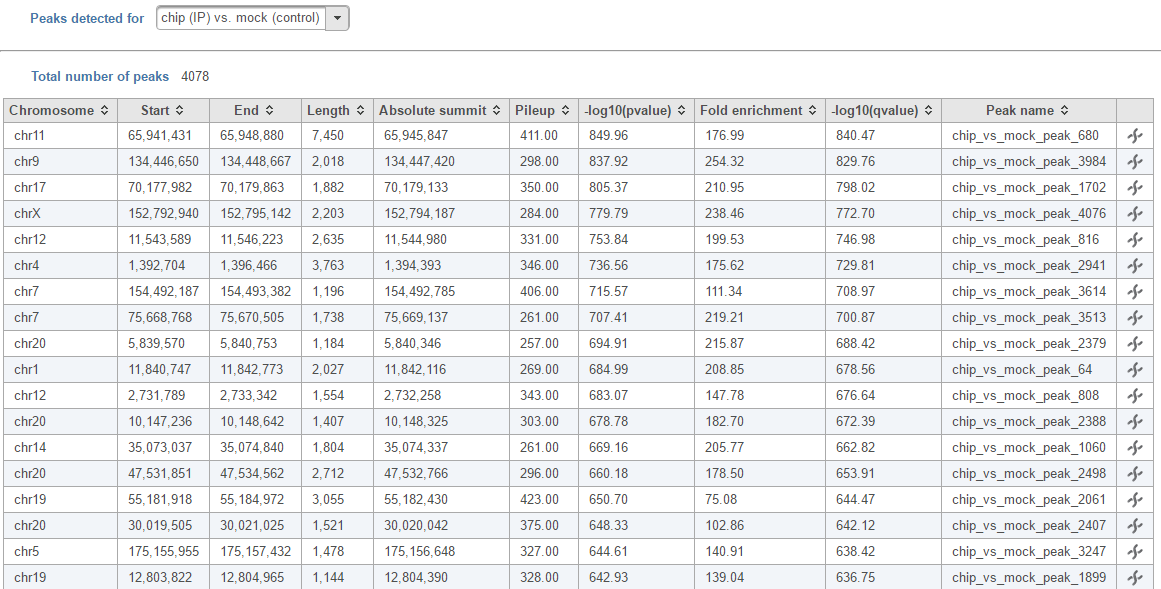
In the task report, each pair will generate a list of peaks displayed in a table (Figure 6)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...