Page History
...
- Click the counts data node
- Click the Exploratory analysis section of the toolbox
- Click Graph-based clustering
- Configure the parameters
- Click Finish to run
Graph-based clustering produces a Clustering result data node. The task report lists the cluster results and cluster statistics (Figure 1). If clustering was run with Split cells by sample enabled on a single cell counts data node, the cluster results table displays the number of clusters found for each sample and clicking the sample name opens the sample-level report.
...
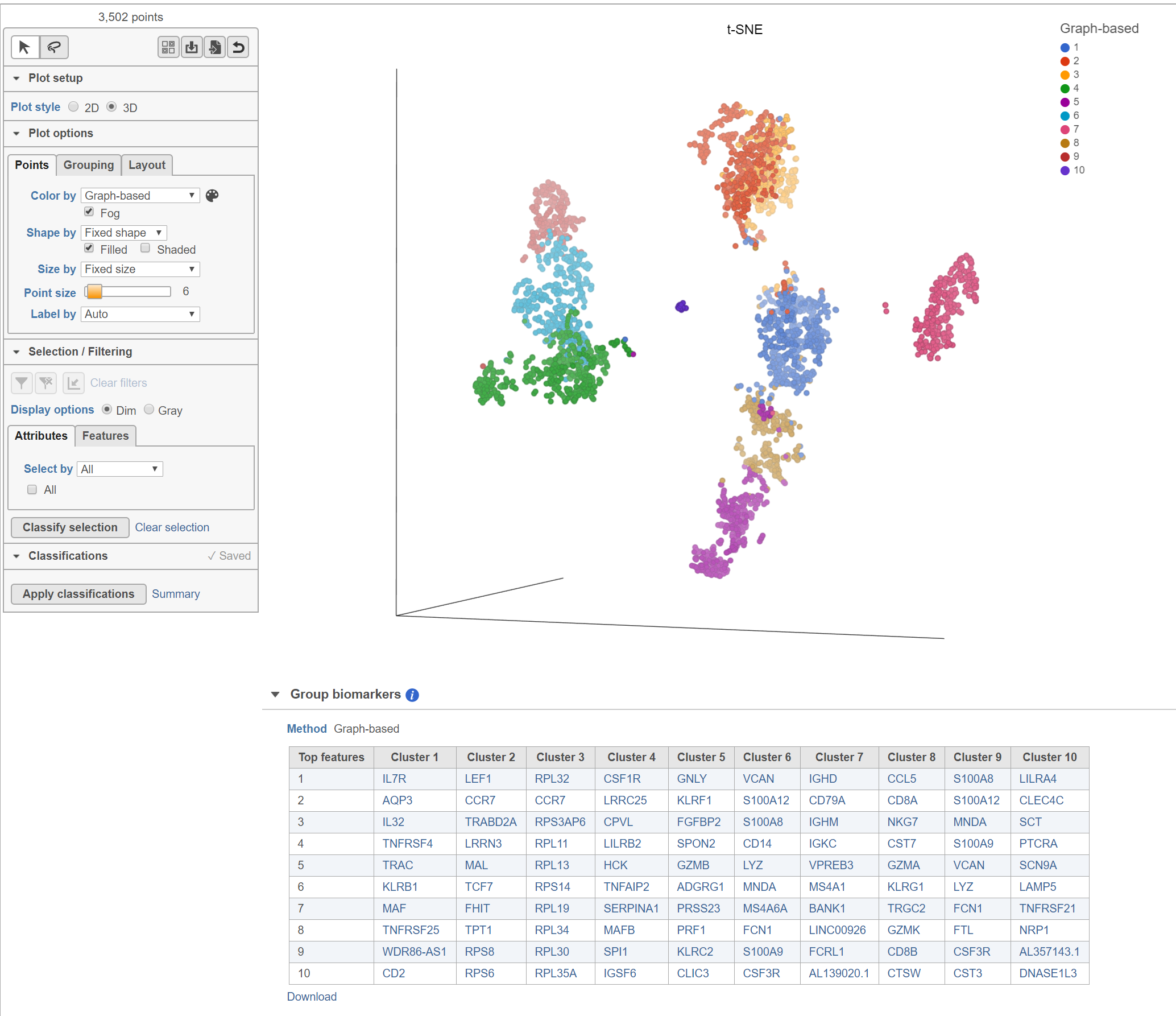
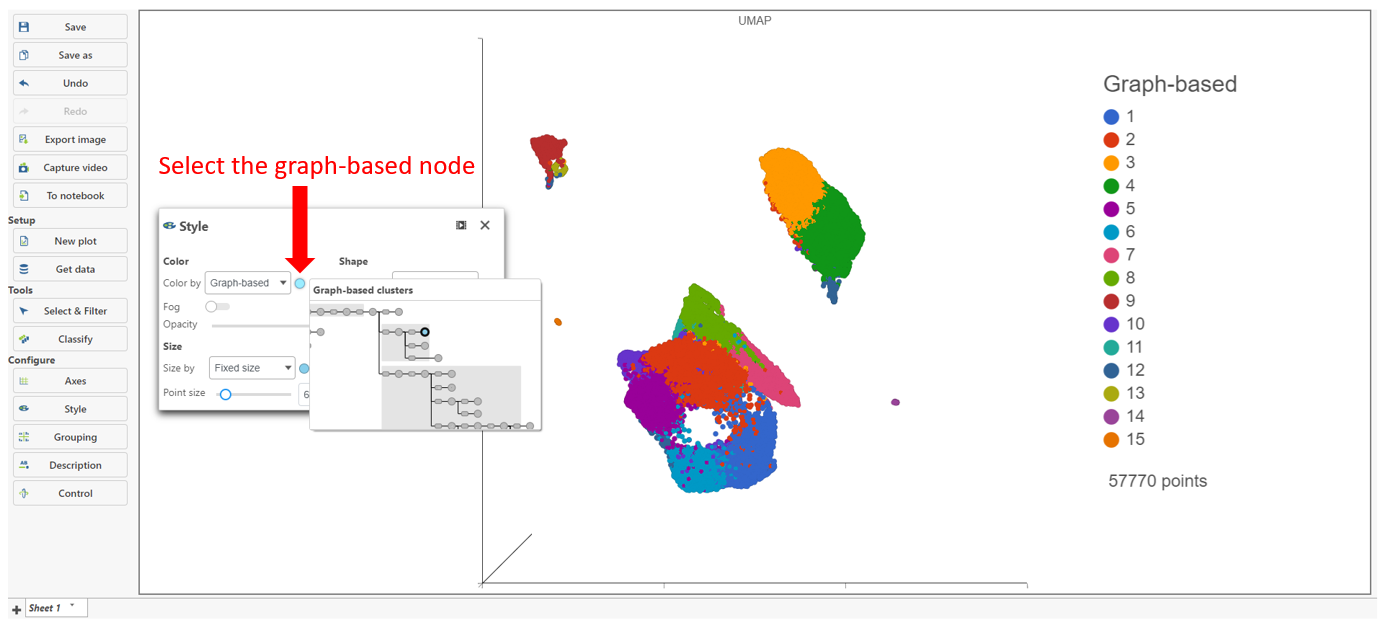
The Clustering result data node includes the input values for each gene and adds cluster assignment as a new attribute, Graph-based, for each observation. If the Clustering result data node is visualized by Scatter plot, PCA, t-SNE, or UMAP, the plot will be colored by the Graph-based attribute and the group biomarker table, if generated, will be included below the plot (Figure 2).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Basic Graph-based clustering parameters
...