Page History
...
Cell Hashing enables sample multiplexing and super-loading in single cell RNA-Seq isolation by labeling each sample with a sample-specific oligo-tagged antibody against a ubiquitously expressed cell surface protein.
The Hashtag demultiplexing task is an implementation of the algorithm used in Stoeckius et al. 20181 for multiplexing cell hashing data. The task adds cell-level attributes "Sample of origin" and " and Cells in droplet".
Prerequisites for running Hashtag demultiplexing
To run Hashtag demultiplexing, your data must meet the following criteria:
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
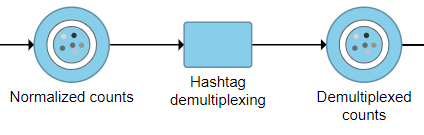
- Click the Normalized counts data node for your cell hashing data
- Click Hashtag demultiplexing in the Pre-analysis tools section of the toolbox
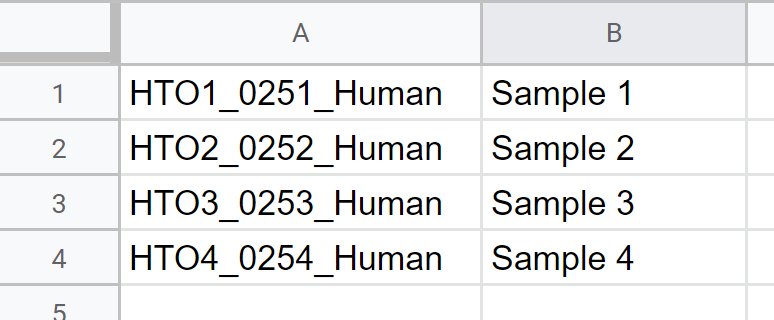
- Click Browse to select your Sample ID file (Optional)
- Click Finish to run
The output is a Dumultiplexed counts data node (Figure 2).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Two cell-level attributes, Cells in droplet and Sample of origin, are added by this task and are available for use in downstream tasks.You can download the attribute values for each cell by clicking the Demultiplexed counts data node, clicking Download, and choosing to download Attributes only.
We recommend using Annotate cells to transfer the new attributes to other sections of your project after downloading the attributes text file.
It is also possible to use the Merge matrices task to combine your data types and attributes.
...