| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Section Heading
TSS plot is available on annotated peak data node. After the peaks are annotated with transcripts, coverage of the reads across the genes overlapped with the regions will be calculated, this information is presented by profile plot (Figure 1) and heatmap (Figure 2).
| Numbered figure captions |
|---|
| SubtitleText | TSS profile plot |
|---|
| AnchorName | tss_profile |
|---|
|
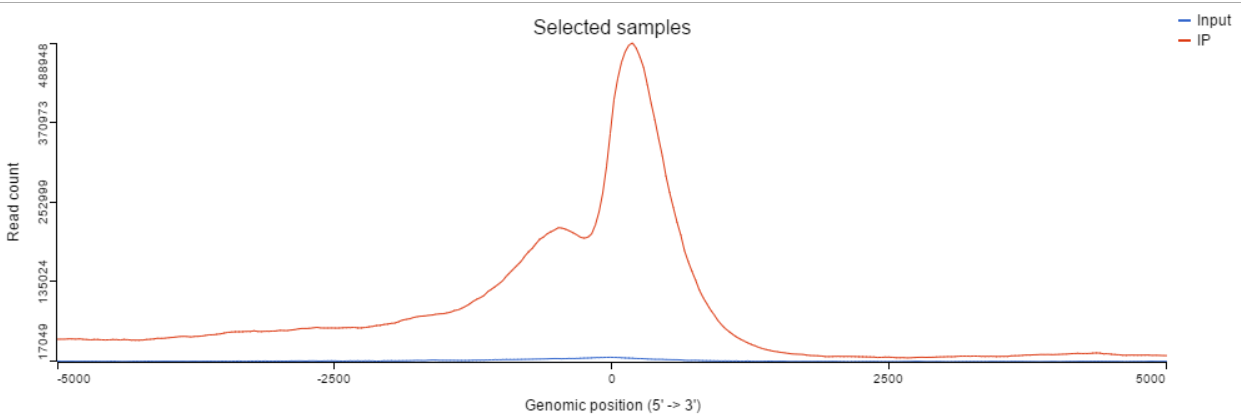 Image Added Image Added
|
TSS profile plot
In the profile plot, the Y-axis is the total read counts, X-axis is the window defined in the annotation task, mid point is the transcription start sites of all the genes overlapped with the peak regions, the start and end point of X-axis is promoter upstream/downstream limit specified in the annotate peaks task (Figure 3). Each line represents a selected sample, all the selected samples are shown in the plot to compare.
By default, all the samples are selected to display, to remove a sample track, click on the  Image Added button next to the sample name on the sample control panel (Figure ); to add a sample in the plot, select the sample name from the Add sample drop-down list, a new line will be added on the plot.
Image Added button next to the sample name on the sample control panel (Figure ); to add a sample in the plot, select the sample name from the Add sample drop-down list, a new line will be added on the plot.
| Numbered figure captions |
|---|
| SubtitleText | Sample selection control panel |
|---|
| AnchorName | select_sample |
|---|
|
 Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | TSS heatmap |
|---|
| AnchorName | tss_heatmap |
|---|
|
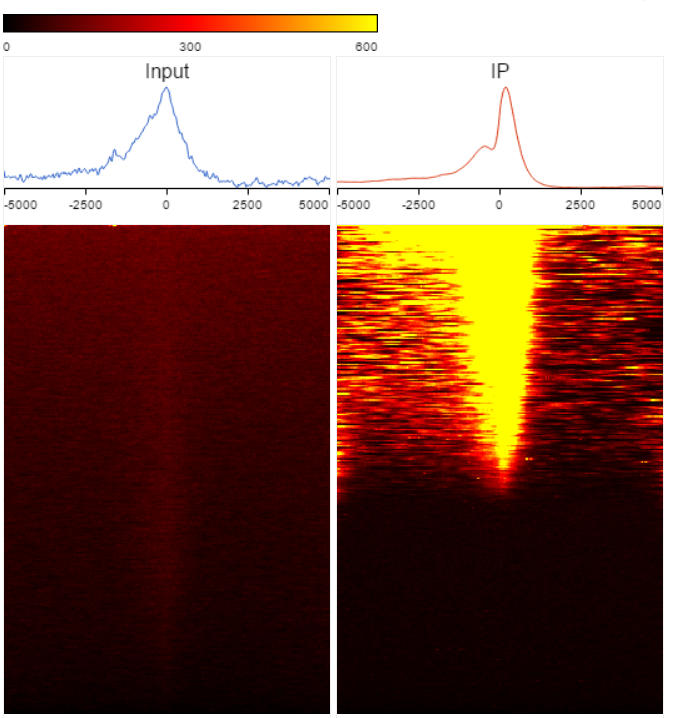 Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | Annotate peaks task dialog |
|---|
| AnchorName | annotation |
|---|
|
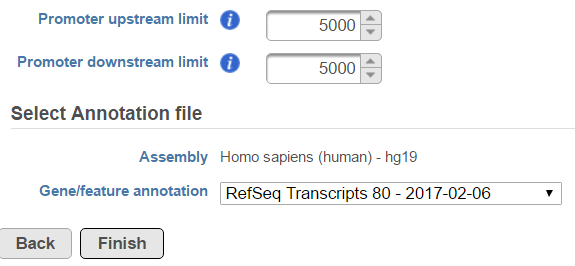 Image Added Image Added
|




