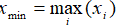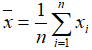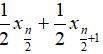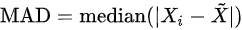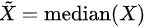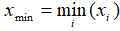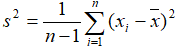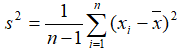...
- Click on a counts data node
- Choose Descriptive Statistics in Pre-analysis tools Statistics section of the toolbox (Figure 1)
| Numbered figure captions |
|---|
| SubtitleText | Descriptive statistics menu |
|---|
| AnchorName | desc |
|---|
|
 Image Removed Image Removed Image Added Image Added
|
This will invoke the dialog (Figure 2), select the calculation configuration dialog; use it to specify which calculation(s) will be performed on samples cells (or cells on a single cell count samples for a bulk analysis data node) or features (Figure 2).
| Numbered figure captions |
|---|
| SubtitleText | Select to calculate descriptive statistics on samples/cells or cellsfeatures |
|---|
| AnchorName | obs |
|---|
|
 Image Removed Image Removed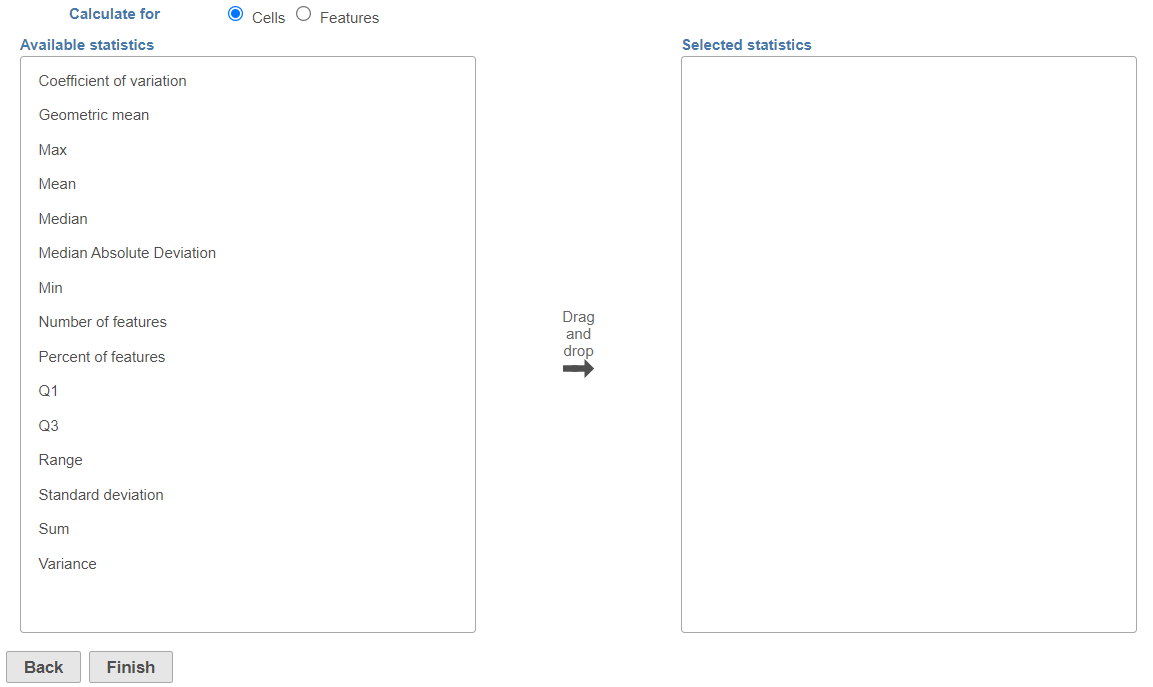 Image Added Image Added
|
The available statistics are listed on the left panel, suppose "x1, x2, ..., xn"represent an array of numbers
- Coefficient of variation (CV):
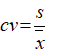 s represent the standard deviation
s represent the standard deviation
- Geometric mean: g=

- Max:
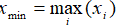
- Mean:
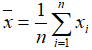
- Median: when n is odd, median is
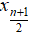 , when n is even, median is
, when n is even, median is 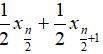
- Median absolute deviation:
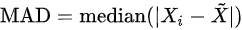 , where
, where 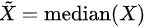
- Min:
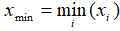
Non zero count: number of observations that is not zero
- Number of cells: Available when Calculate for is set to Features. Reports the number of cells with the value [<, <=, =, !=, > >=] (select one from the drop down list) than the cut off value entered in the text box. The cut off will be applied to the values present in the input data node, i.e. if invoked on non-normalised data node, the values are raw counts. For instance, use this option if you want to know the number of cells in which each feature was detected; possible filter: Number of cells whose value > 0.0
- Percent of cells: Available when Calculate for is set to Features. Reports the number of cells with the value [<, <=, =, !=, > >=] (select one from the drop down list) than the cut off value entered in the text box.
- Number of features: Available when Calculate for is set to Cells. Reports the number of features with the value [<, <=, =, !=, > >=] (select one from the drop down list) than the cut off value entered in the text box. The cut off will be applied to the values present in the input data node, i.e. if invoked on non-normalised data node, the values are raw counts. For example, use this option if you want to know the number of detected genes per each cell; filter: Number of features whose value > 0.0
- Percent of features: Available when Calculate for is set to Cells. Reports the fraction of features with the value [<, <=, =, !=, > >=] (select one from the drop down list) than the cut off value entered in the text box.
- Q1: 25th percentile
- Q3: 75th percentile
- Range: xmax - x min
- Standard deviation:
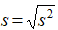 where
where 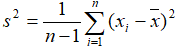
- Sum:
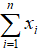
- Variance:
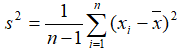
...
The output of the task is a matrix, it : Cell stats (result of Calculate for Cells) or Feature stats (result of Calculate for Features) (Figure 4). The results can be visualized in the Data Viewer.
| Numbered figure captions |
|---|
| SubtitleText | Descriptive statistics task produces either a Cell stats (calculation per cell) or Feature stats (calculation per feature) data node |
|---|
| AnchorName | stats-nodes |
|---|
|
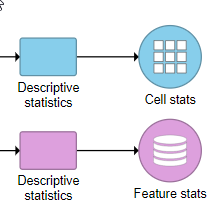 Image Added Image Added
|
...