Page History
...
Now that the data has been imported, we need to make a few changes to the data annotation before analysis.
Modifying
...
sample attributes
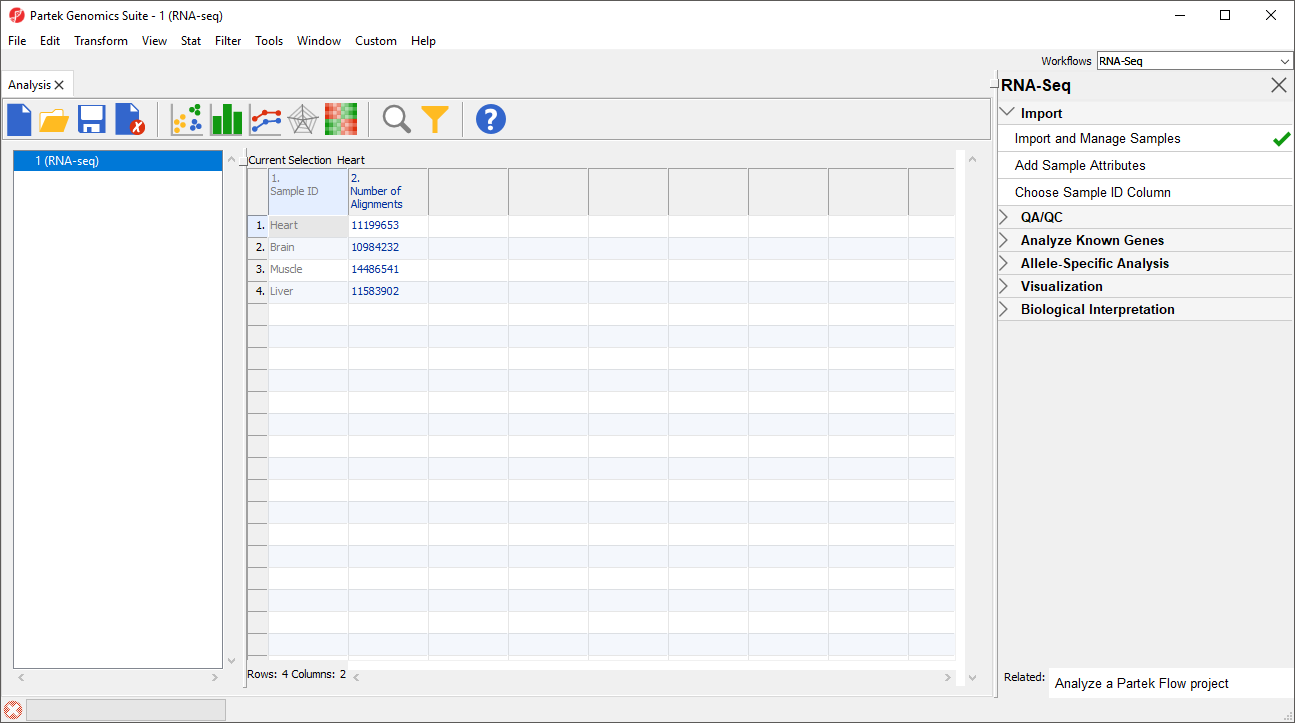
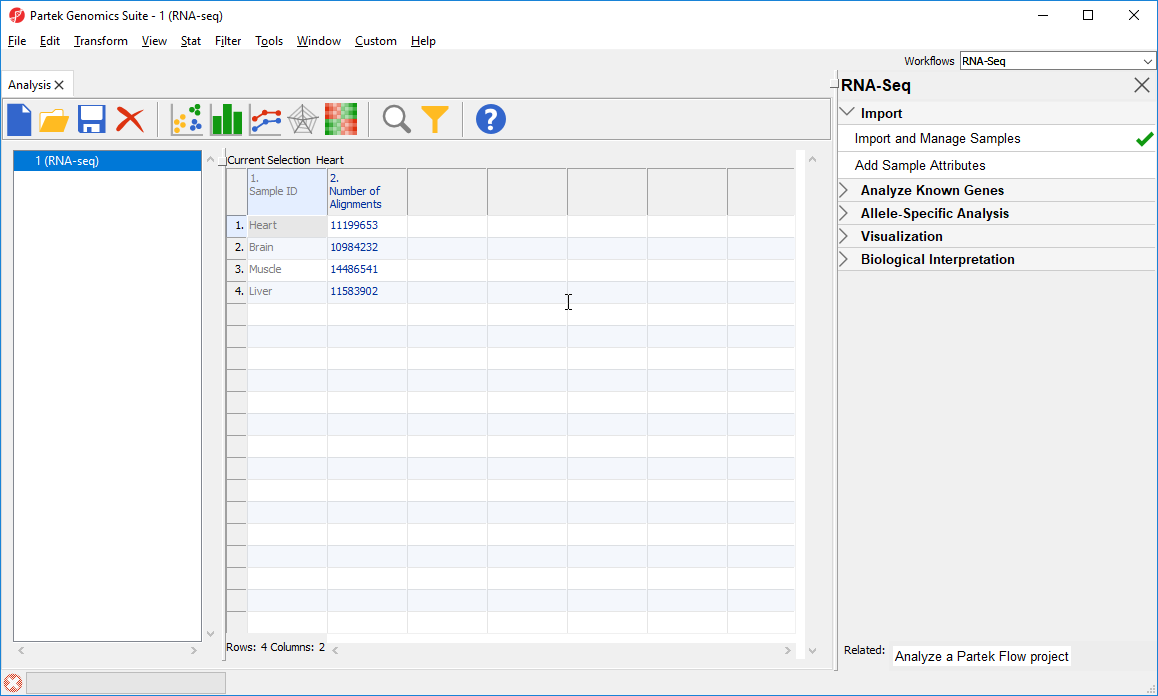
Notice that the Sample ID names in column 1 are gray (Figure 1). This indicates that Sample ID is a text factor. Text factors cannot be used as a variable in downstream analysis so we need to change Sample ID to a categorical factor.
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
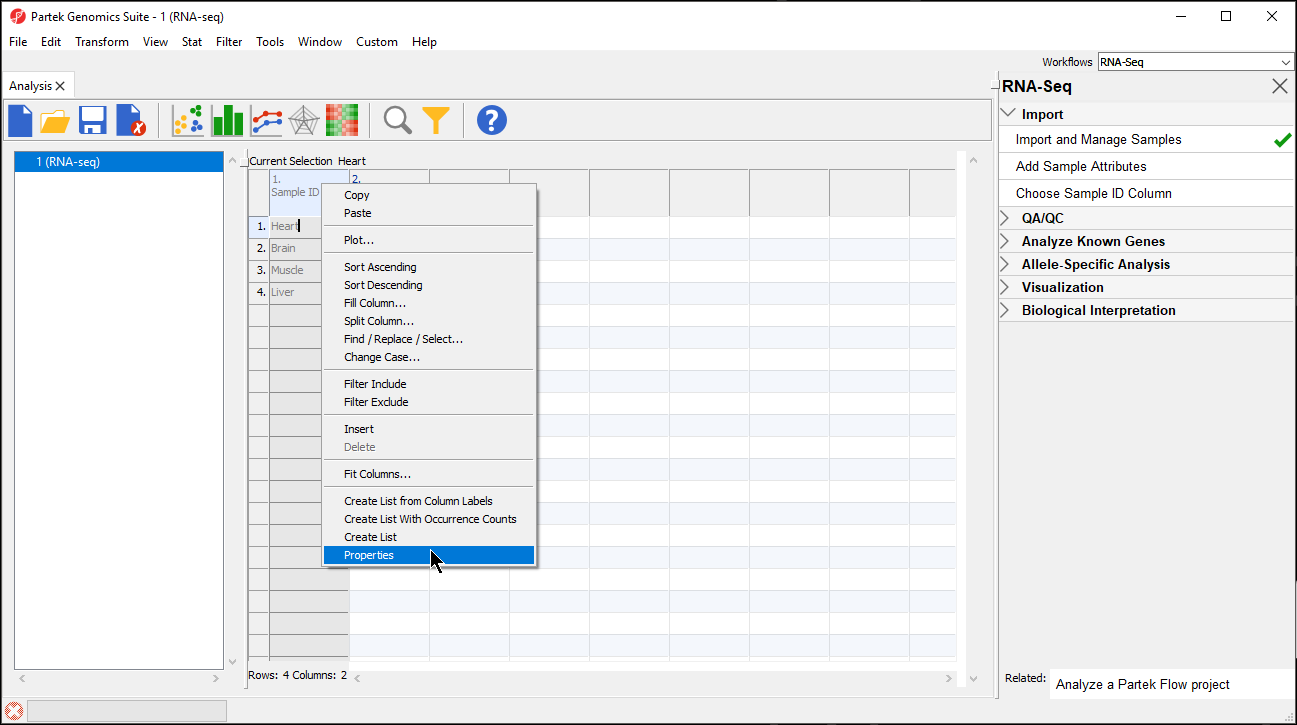
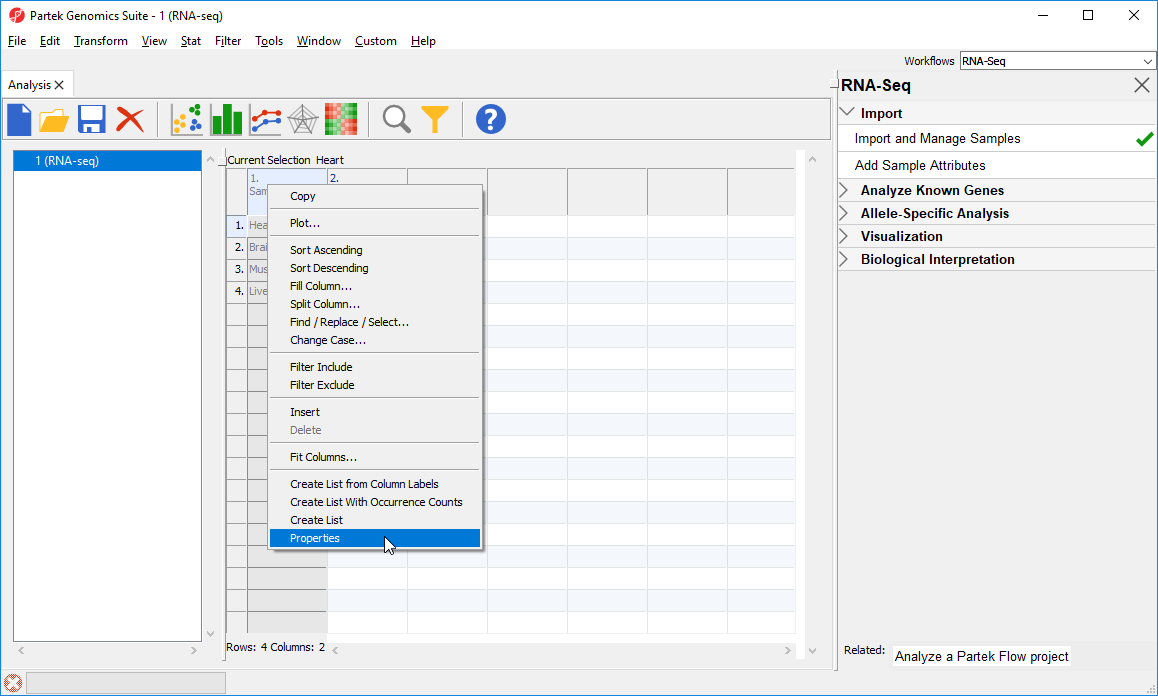
- Right-click on the column header to invoke the contextual menu and then select pop-up menu
- Select Properties (Figure 2)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
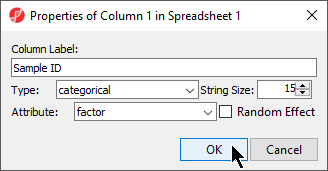
- Configure the Properties of Column 1 in Spreadsheet 1 dialog as shown (Figure 3) with Type set to categorical and Attribute set to factor
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select OK
The samples names in column 1 are now black, indicating that they have been changed to a categorical variable. Next, we will add attributes for grouping the data.
Adding
...
sample attributes
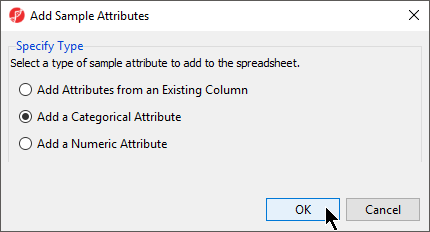
- From the RNA-seq workflow panel, select Add sample attributeSample Attributes to bring up the Add Sample Attributes dialog (Figure 4)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select the Select Add a categorical attribute optionCategorical Attribute
- Select OK to bring up the Create categorical attribute dialog
...
The attribute will now appear as a new column in the RNA-seq spreadsheet with the heading Tissue and the groups muscle and not muscle.
Choosing Sample ID
...
column
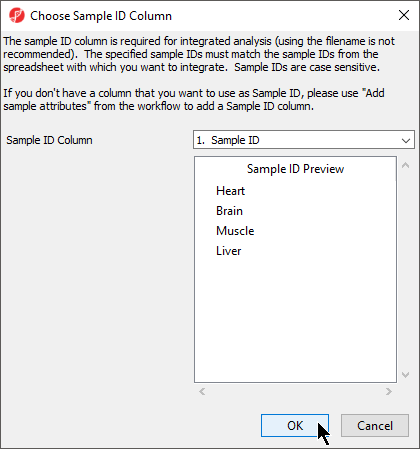
The next available step in the Import panel of the RNA-seq workflow is Choose Sample ID Column. Verifying the correct column is designated the Sample ID becomes particularly important when data from multiple experiments is being combined.
- Select Choose Sample ID Column from the Import panel of the RNA-seq Seq workflow
- Select OK (Figure 6)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
| Page Turner | ||
|---|---|---|
|
...