| Table of Contents |
|---|
| exclude | Additional Assistance |
|---|
|
What is
...
Cell Ranger?
Enrichment analysis is a technique commonly used to add biological context to a list of genes, such as list of significant genes. The procedure is based on assigning genes to groups and then finding overrepresented groups in filtered gene lists using a Fisher's exact test.
Running Gene set enrichment
We recommend filtering to a set of genes you want to test for enrichment, but Gene set enrichment will run on any Feature list data node.
- Click a Feature list data node
- Click the Biological interpretation section of the toolbox
- Click Gene set enrichment
- Configure the background gene list (optional)
The background gene list is used as the list of possible genes. By default, this is the genes included in the selected gene set database. If your assay limits the genes that could be detected, you may want to specify a background list.
The gene sets available for the current Assembly are listed under the Gene set drop-down list. The assembly is automatically selected, if possible. If the assembly cannot be detected, you can specify it using a drop-down menu.
...
Cell Ranger is a set of analysis pipelines that process Chromium single cell data to align reads, generate feature-barcode matrices and perform clustering and gene expression analysis for 10X Genomics Chromium Technology[1].
Cell Ranger - Gene Expression in Partek Flow
The 'cellranger count' pipeline from Cell Ranger v6.0.0[2] has been wrapped in Partek Flow as Cell Ranger - Gene Expression task. It does not comprehensively cover all of the options and analysis cases Cell Ranger can handle for now, but converts FASTQ files from 'cellranger mkfastq' and performs alignment, filtering, barcode counting, UMI counting for single cell gene expression and Feature Barcode data. The output gene expression count matrix in .h5 format (both raw and filtered available for users to download in the output page of task details) then becomes the starting point for downstream analysis for scRNA-seq in Flow. For Feature Barcode data, Flow outputs a unified feature-barcode matrix that contains gene expression counts alongside Feature Barcode counts for each cell barcode.
Note: When use Cell Ranger - Gene Expression task in Partek Flow, there are more restrictions on sample name -- sample name can only contain letters, digits, underscores and dashes. Please edit the sample names on Data tab in Partek Flow to remove any other characters ,e.g. space etc.
Running Cell Ranger - Gene Expression in Flow
To run the Cell Ranger - Gene Expression task for scRNA-seq data in Flow, select Unaligned reads datanode, then select Cell Ranger - Gene Expression in the 10x Genomics section (left panel, Figure 1). For Feature Barcode data, there will be two data nodes once the FASTQ files have been imported into Flow properly - mRNA and protein (right panel, Figure 1). Users should select mRNA datanode to trigger the Cell ranger - Gene Expression task.
| Numbered figure captions |
|---|
| SubtitleText | Selecting the gene set for Enrichment analysis. Sets available for the current Assembly are listed under Gene set |
|---|
| AnchorName | Gene set selection |
|---|
|
 Image Removed Image Removed
|
By default, the groups are defined by Gene Ontology (GO), a bioinformatics initiative to unify the representation of gene and gene product attributes across various species [1, 2].
...
| Cell Ranger - Gene Expression task for converting fastqs to Single cell counts. | | AnchorName | Task selection |
|---|
|
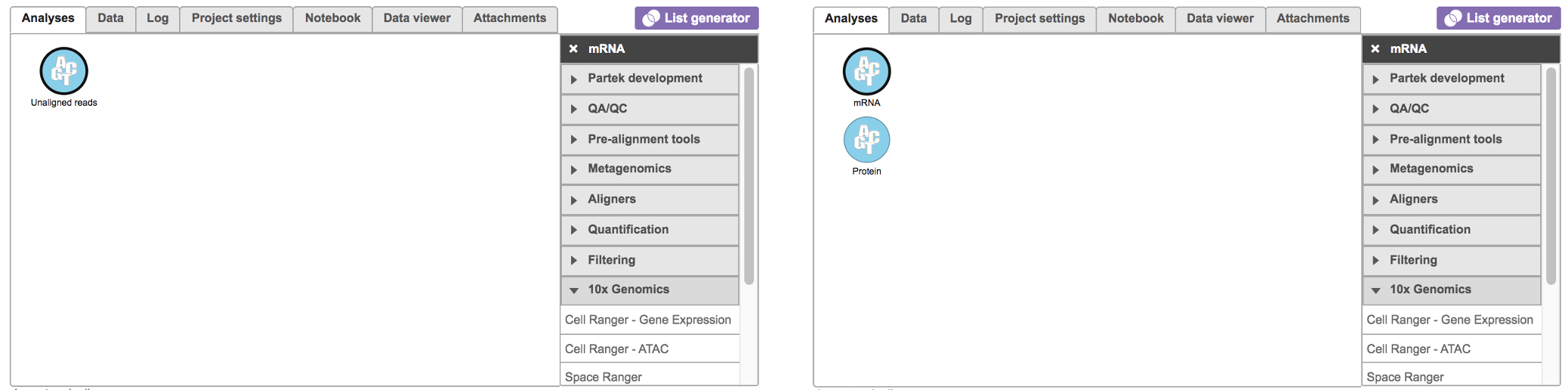 Image Added Image Added
|
Once the Genomics production has been picked, users will be asked to create a Reference assembly if it is the first time to run the Cell Ranger - Gene Expression task (Figure 2). In Partek Flow, we will use Cell Ranger ARC 2.0.0 to create reference assembly for all 10x Genomics analysis pipelines. To create and use a reference assembly, Cell Ranger ARC requires a reference genome sequence (FASTA file) and gene annotations (GTF file), here are the details.
| Numbered figure captions |
|---|
| SubtitleText | Create Cell Ranger ARC reference genome for the first time user. |
|---|
| AnchorName | Create reference |
|---|
|
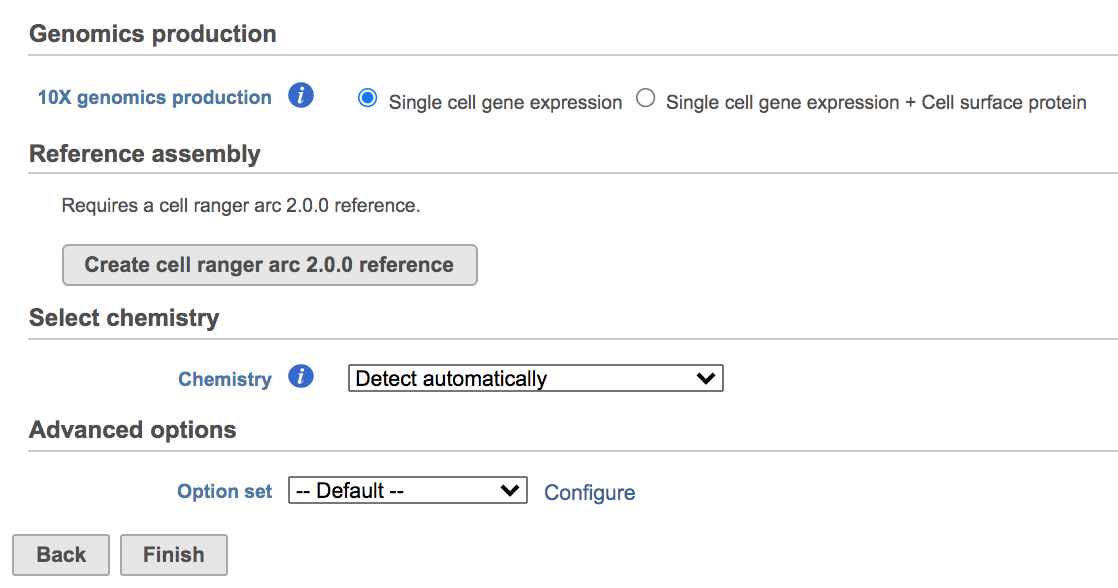 Image Added Image Added
|
Clicking the big grey button of Create Cell Ranger ARC 2.0.0 reference would pop up a new window where lists the requirements that users need to fill in (Figure 3). To create the same reference genomes (2020-A) that are provided in Cell Ranger by default, the transcriptome annotations are respectively GENCODE v32 for human and vM23 for mouse, which are equivalent to Ensembl 98[3]. If users don't have any options in the dropdown list, they can click Add annotation model (GTF file) for Index, or New assembly... (FASTA file)for Assembly and upload the files.
| Numbered figure captions |
|---|
| SubtitleText | Adding gene set files via Create gene list dialog. Download gene set obtains a gene set file from Partek (human, mouse and rat are supported), Import gene set opens a file browser, which is used to specify the file that should be added to the Library file management functionality |
|---|
| AnchorName | Create gene list dialog |
|---|
|
 Image Removed Image Removed
|
The result is stored under an Enrichment task node. To open it, double click on the node or select the respective Task report from the context sensitive menu.
Gene set enrichment task report
...
| Build a new reference assembly with Flow Cell Ranger ARC. | | AnchorName | Create reference |
|---|
|
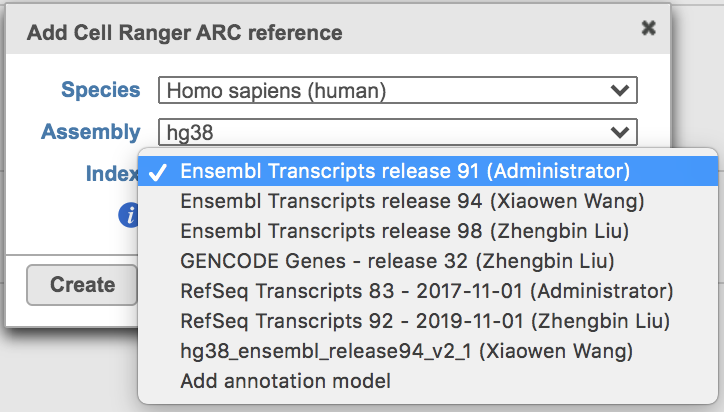 Image Added Image Added
|
Once the right options has been chosen/provided, simply press the Create button to finish. The reference assembly of ‘Homo sapiens (human) - hg38’ has been created as an example here (Figure 4).
| Numbered figure captions |
|---|
| SubtitleText | Create Human reference genome (hg38) for Cell Ranger - Gene Expression. |
|---|
| AnchorName | Create reference |
|---|
|
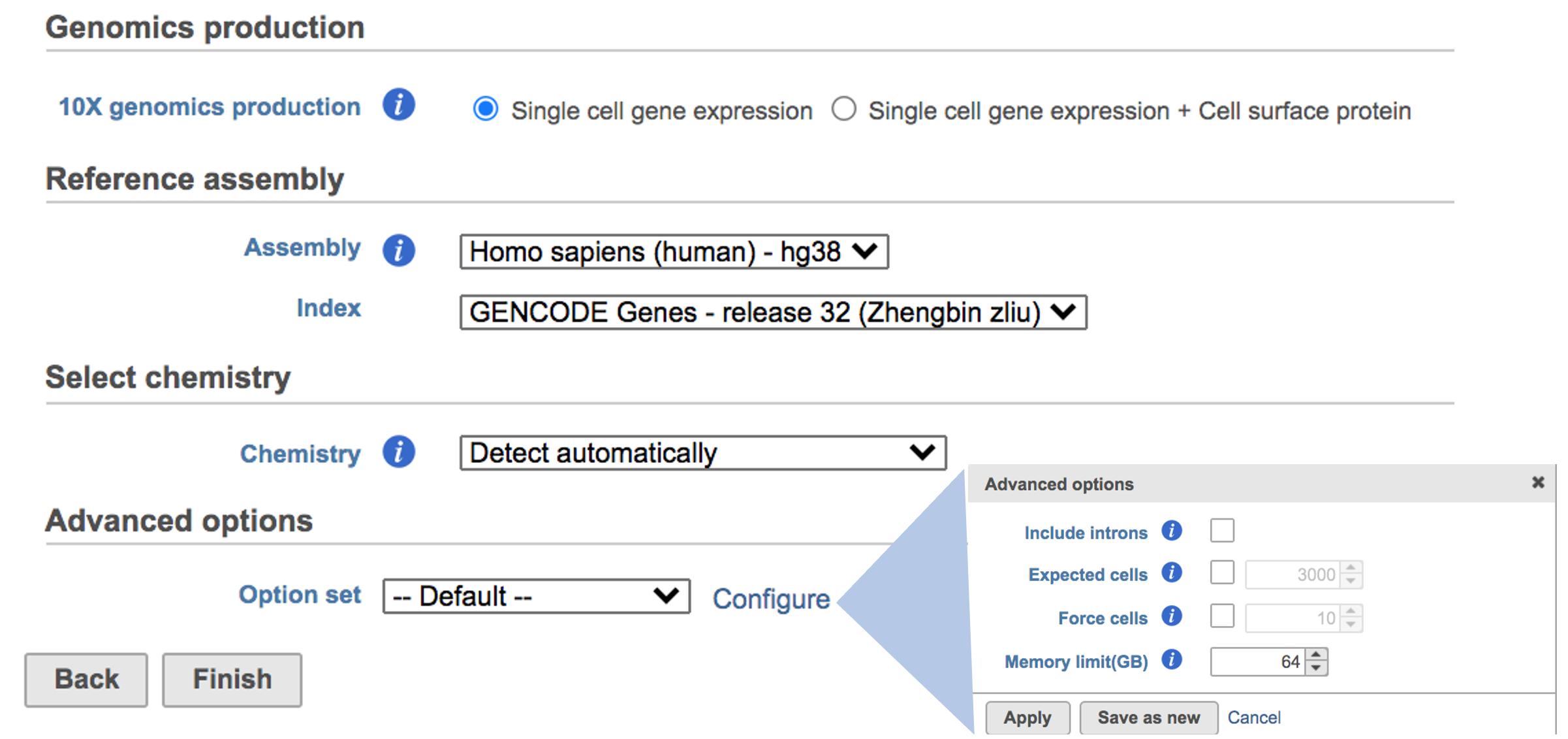 Image Added Image Added
|
The main task menu will be refreshed as above (Figure 4) for gene expression data if references have been added. Users can go ahead click the Finish button to run the task as default.
While for Feature Barcode data, there are more information needed besides reference assembly. An additional section of Protein has been added to the interface if Single cell gene expression + Cell surface protein has been selected for Feature Barcode data (Figure 5). Users need firstly push the button Select data node and select the correct data for feature of antibody capture or protein in a new pop-up window (top right, Figure 5). Then users need to upload the feature reference file (.csv) prepared for their datasets. A Feature Reference CSV file declares the molecule structure and unique Feature Barcode sequence of each feature present in the experiment. It should include at least six columns: id, name, read, pattern, sequence and feature_type. An example of TotalSeq™-B Feature Reference CSV has been linked here. Users can download it by clicking the link and use it as a template for their own data. But for more details, please refer to 10x Genomics webpage[4].
| Numbered figure captions |
|---|
| SubtitleText | Go enrichment report (truncated). Gene set column contains Gene Ontology identifiers (hyperlinks). Category labels are in the Description column. Enrichment score: negative natural logarithm of the enrichment P-value derived from the Fisher's exact test. Genes in list: number of genes that are present both in the list of significant genes and the gene set (GO category). Genes not in list: number of genes that are present in the gene set, but are not present in the list of significant genes. The column on the right contains links to gene breakdown chart and extra details |
|---|
| AnchorName | Go enrichment report |
|---|
|
 Image Removed Image Removed
|
...
| Run Cell Ranger - Gene Expression task for Feature Barcode data with reference assembly hg38 in Flow. | | AnchorName | Run Cell ranger task |
|---|
|
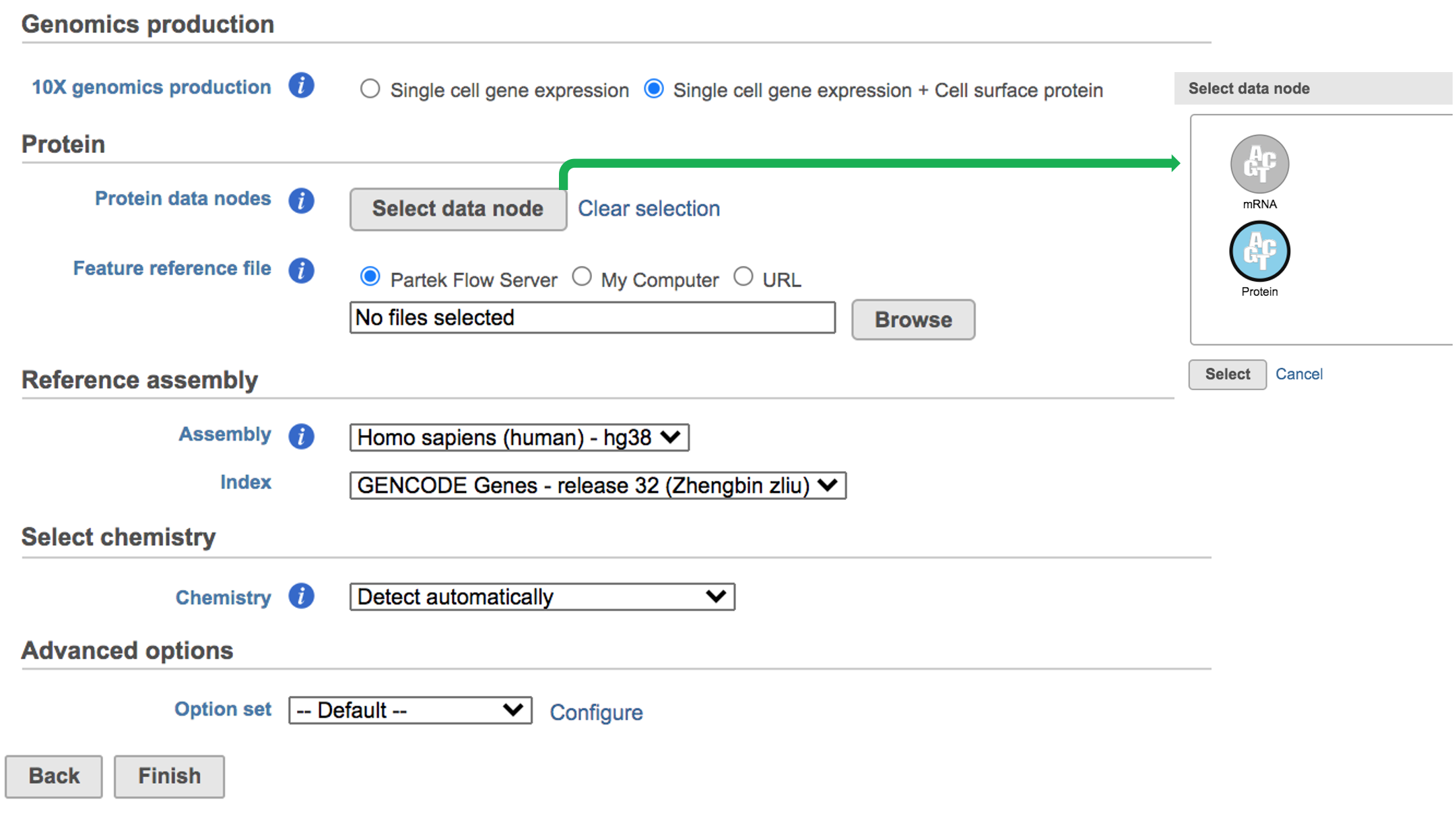 Image Added Image Added
|
A new data node named Single cell counts will be displayed in Flow if the task has been finished successfully (Figure 6). This data node contains a filtered feature barcode count matrix for gene expression data, but a unified feature-barcode matrix that contains gene expression counts alongside Feature Barcode counts for each cell barcode for Feature Barcode data. To open the task report when the task is finished, double click the output data node, or select the Task report in the Task results section after single clicking the data node. Users then will find the task report (Figure 7) is the same to the ‘Summary HTML’ from Cell Ranger output.
| Numbered figure captions |
|---|
| SubtitleText | The finished Cell Ranger - Gene Expression task in Flow. Feature Barcode data has been used in the example task here. |
|---|
| AnchorName | Run Cell ranger task |
|---|
|
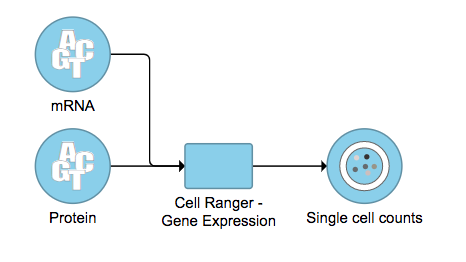 Image Added Image Added
|
Cell Ranger - Gene Expression task report in Flow
Task report is sample based. Users can use the dropdown list on the top left to switch samples. Under the sample name, there are two tabs on each report - Summary report and Analysis report (Figure 7). Important information on Estimated Number of Cells, Mean Reads per Cell, Median Genes per Cell, as well as information on Sequencing, Mapping, and Sample are summarized in different panels. The Barcode Rank Plot has also been included as an important piece in the Cells panel in the Summary report (Figure 7).
| Numbered figure captions |
|---|
| SubtitleText | Contingency table used to calculate the enrichment p-value. List refers to the list of significant genes, set refers to the gene ontology category |
|---|
| AnchorName | Contingency table |
|---|
|
 Image Removed Image Removed
|
...
| The example report of Cell ranger task in Flow. | | AnchorName | Cell ranger task report |
|---|
|
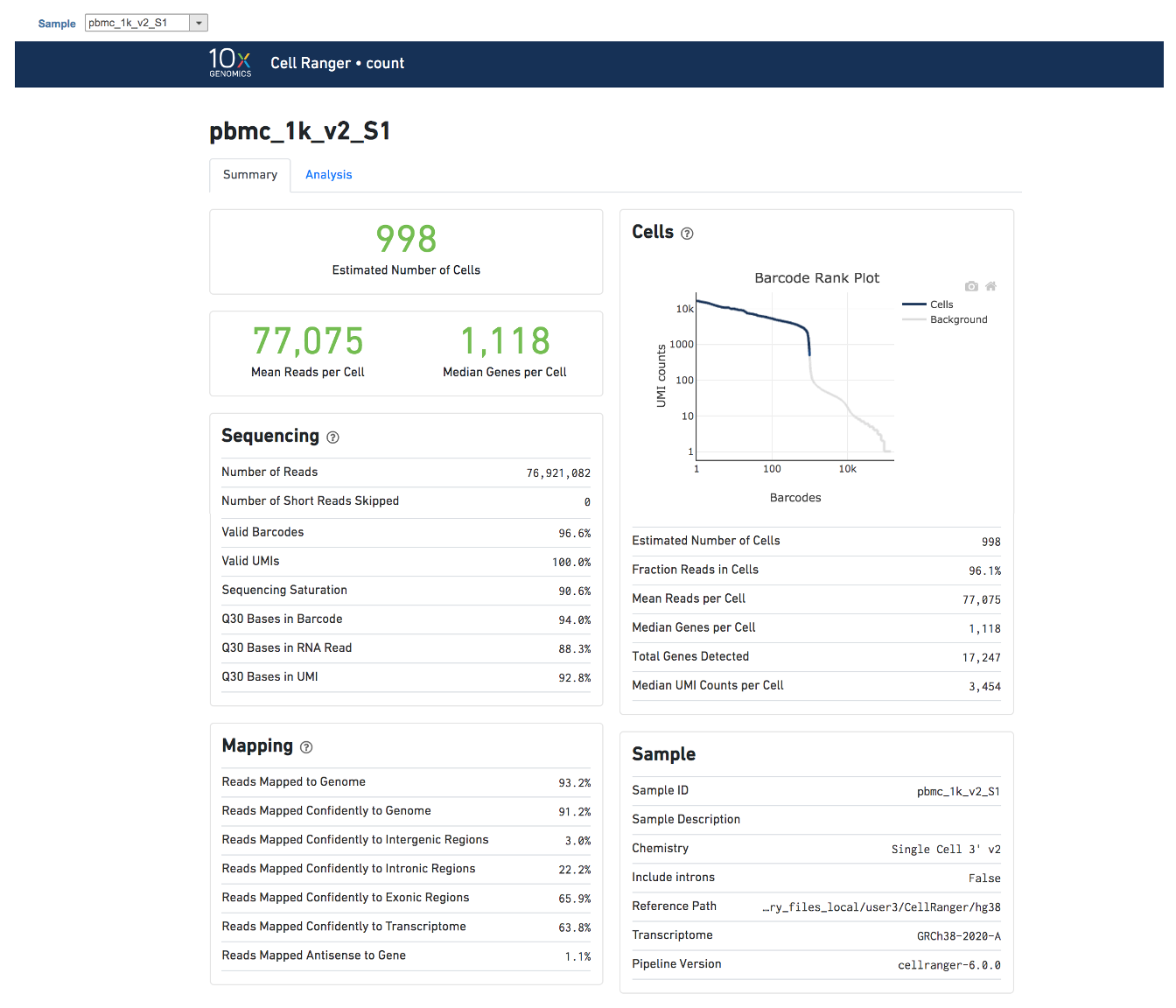 Image Added Image Added
|
Another two plots -biplots of Sequencing Saturation and Median Genes per Cell to Mean Reads per Cell have been included in the Analysis report as they are important metrics to library complexity and sequencing depth (Figure 8).
| Numbered figure captions |
|---|
| SubtitleText | Gene ontology enrichment extra details Analysis report of Cell ranger task in Flow. |
|---|
| AnchorName | Extra enrichment details |
|---|
|
 Image Removed Image Removed
|
...
|
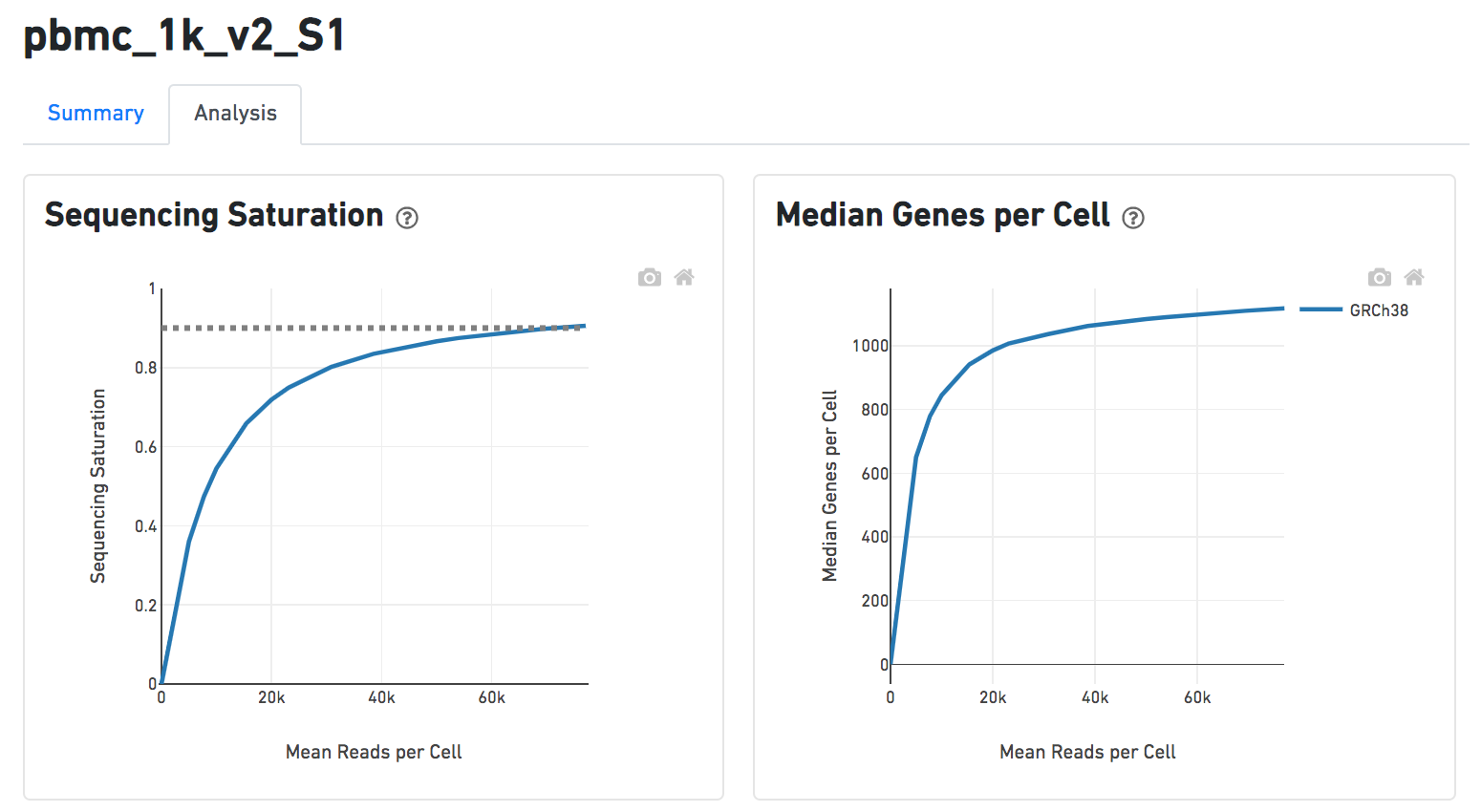 Image Added Image Added
|
Details will be exhibited and the panel will be expanded correspondingly if the the  Image Added icon is clicked. In the example below, the plot of Median Genes per Cell has been expanded while the Sequencing Saturation plot hasn't (Figure 9).
Image Added icon is clicked. In the example below, the plot of Median Genes per Cell has been expanded while the Sequencing Saturation plot hasn't (Figure 9).
| Numbered figure captions |
|---|
| SubtitleText | Selecting a GO category in the table report opens up a browser and displays additional information on that category via GO web-page |
|---|
| AnchorName | GO category selection |
|---|
|
 Image Removed Image Removed
|
References
...
| Expanded panel of Cell ranger task report in Flow. | | AnchorName | Cell ranger task report |
|---|
|
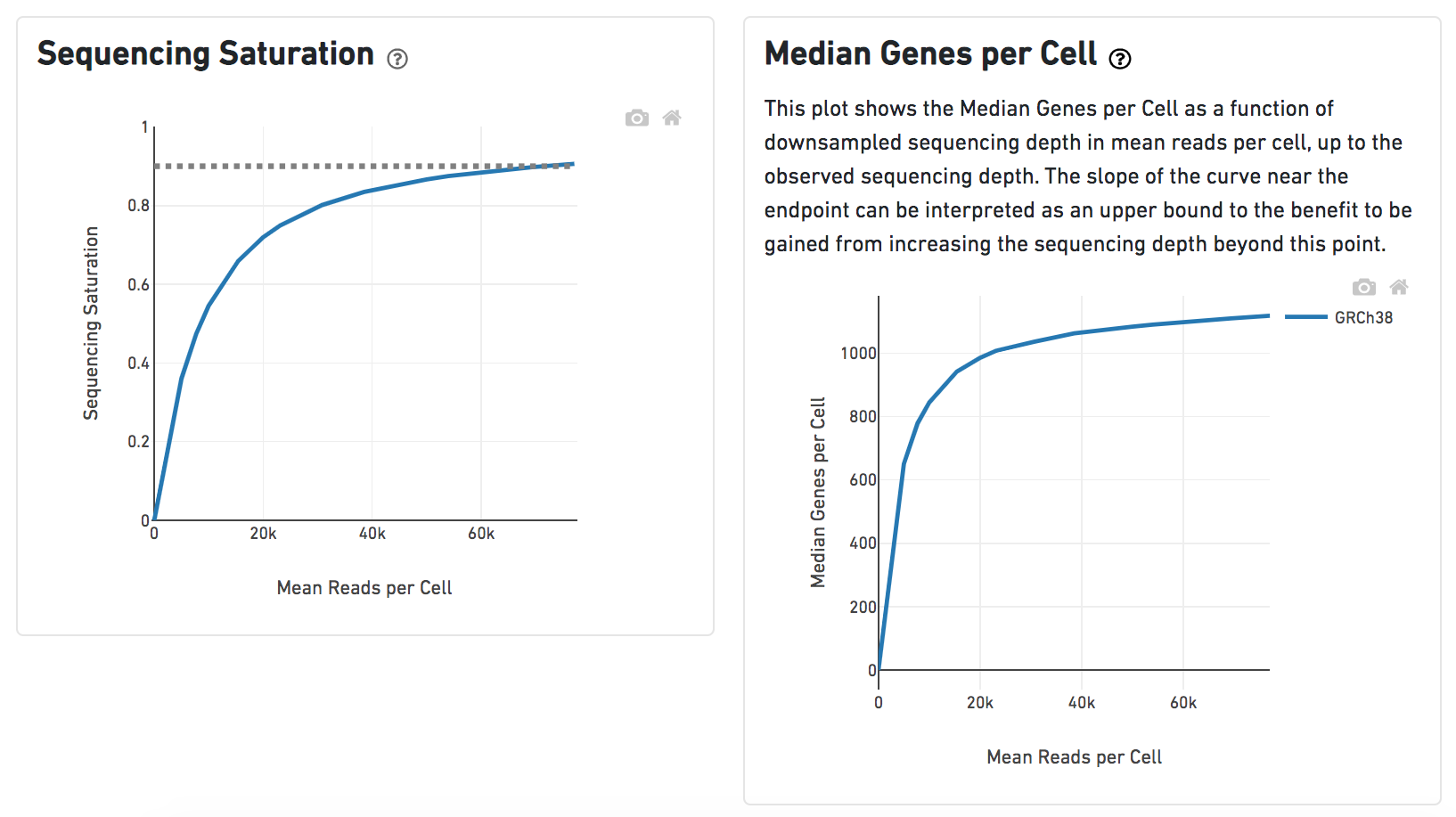 Image Added Image Added
|
Other than two additional panels summarized information for Antibody Sequencing and Antibody Application have been added, the task report for Feature Barcode data is the same to scRNA-seq data report.
| Numbered figure captions |
|---|
| SubtitleText | Additional panels of Cell Ranger - Gene Expression task report for Feature Barcode data in Flow. |
|---|
| AnchorName | Cell ranger task report |
|---|
|
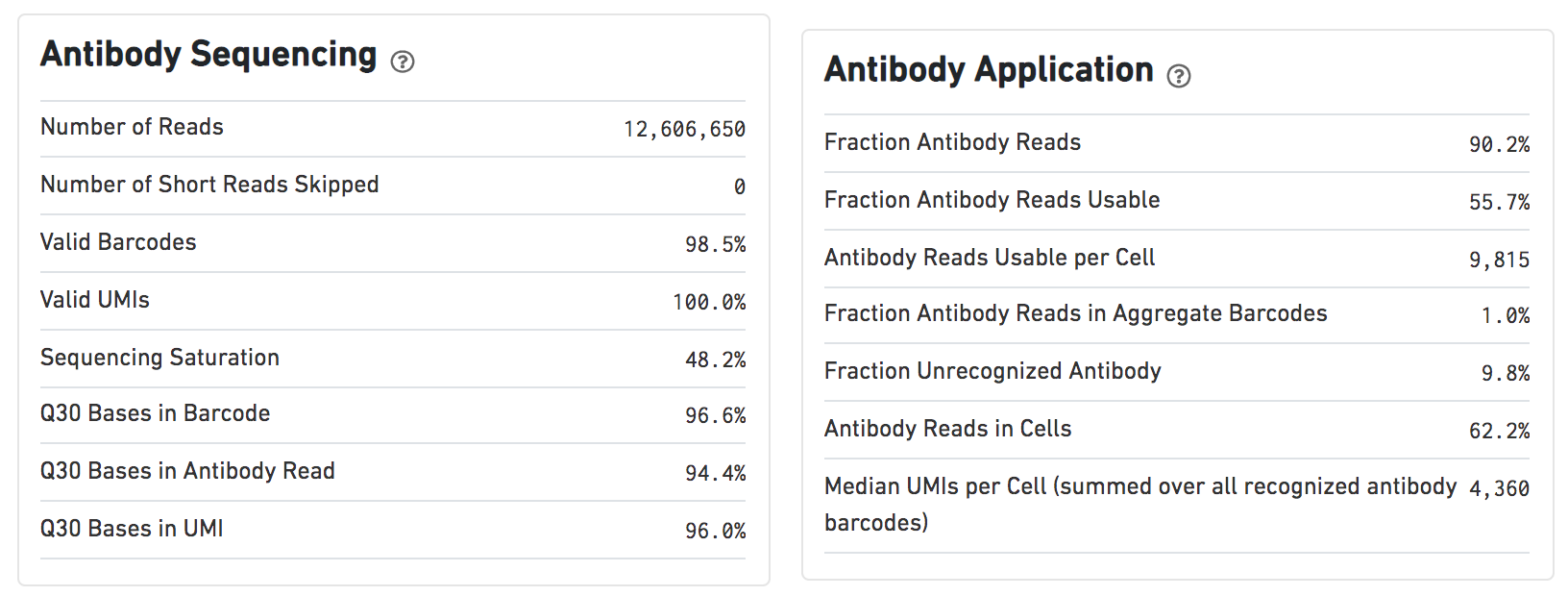 Image Added Image Added
|
Users can click Configure to change the default settings in Advanced options (Figure 4).
Include introns: Count reads mapping to intronic regions. This may improve sensitivity for samples with a significant amount of pre-mRNA molecules, such as nuclei.
Expected cells: Expected number of recovered cells. Default: 3,000 cells.
Force cells: Force pipeline to use this number of cells, bypassing the cell detection algorithm. Use this if the number of cells estimated by Cell Ranger is not consistent with the barcode rank plot.
Memory limit (GB): Restricts Cell Ranger - Gene Expression to use specified amount of memory (in GB) to execute pipeline stages.
References
https://support.10xgenomics.com/single-cell-gene-expression/software/overview/welcome
https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/6.0/release-notes
- https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/4.0/release-notes
- https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/latest/using/feature-bc-analysis#feature-ref
...














