Page History
A fusion gene is a hybrid gene that combines parts of two or more original genes. They can form as a result of chromosomal rearrangements (such as translocation, interstitial deletion, or chromosomal inversion) or abnormal transcription and have been shown to act as drivers of malignant transformation or/and progression in various neoplasms (1). The discovery and characterization of fusion genes hav have been greatly facilitated by the use of NGS (2) and several computational algorithms have been developed to detect them.
...
| Table of Contents | ||||||
|---|---|---|---|---|---|---|
|
...
STAR Algorithm
General Overview
The Partek® Flow® fusion detection algorithm uses paired-end information to find pairs of genes that may express as a hybrid. A paired-end read is considered for a fusion event if:
- an alignment from the first-in-pair maps to a different sequence (chromosome) than an alignment from the second-in-pair, or;
- the distance between all alignments from the first-in-pair and the second-in-pair exceed a custom-defined threshold (default: 50 kb).
The algorithm then reports peaks of reads that are potentially involved in a fusion event. Adjacent peaks are merged if their distance is less than 200 bp (default) and the probability that the peak is derived from the null distribution of peaks (determined by permutation) is reported. False positives hits are reduced by ignoring alignments that overlap with regions masked in the .2bit file. Finally, the peaks are annotated with a transcript model and a report is generated for pairs of peaks which map to different transcripts.
Running Partek Fusion Gene Algorithm within Partek Flow
Partek algorithm can be invoked on a data node containing aligned paired-end reads (i.e. Aligned reads node), through the Detect fusion genes link in the Variant detection section of the toolbox (Figure 1).
The STAR aligner also has the ability to detect fusion genes (referred to as “chimeric alignments”) (5,6). During the first phase of alignment, STAR searches for maximal mappable prefixes (seeds) of sequencing reads. In the second phase, all the seeds that align within user-defined genomic windows are stitched together. If an alignment within one genomic window does not cover the entire read sequence, STAR will try to find two or more windows that cover the entire read. This essentially results in the detection of fusion events, with different parts of reads aligning to distal genomic locations, or different chromosomes, or different strands.
STAR fusion detection is performed in two steps: chimeric alignment of reads with the STAR aligner and fusion detection with STAR-Fusion. Performing fusion detection in two steps is equivalent to running the analysis in "Kickstart" mode, as described by the authors of STAR-Fusion. We recommend using STAR version 2.7.8a (see Task management to check which version you are running).
To save time, you can import the pre-built STAR-Fusion pipeline from our hosted pipeline page. This pipeline includes the two steps outlined below, where the advanced options for the STAR 2.7.8a alignment have been optimized for fusion detection according to the STAR-Fusion author's recommendations. See Importing a Pipeline for more information.
Running STAR Chimeric Alignment within Partek Flow
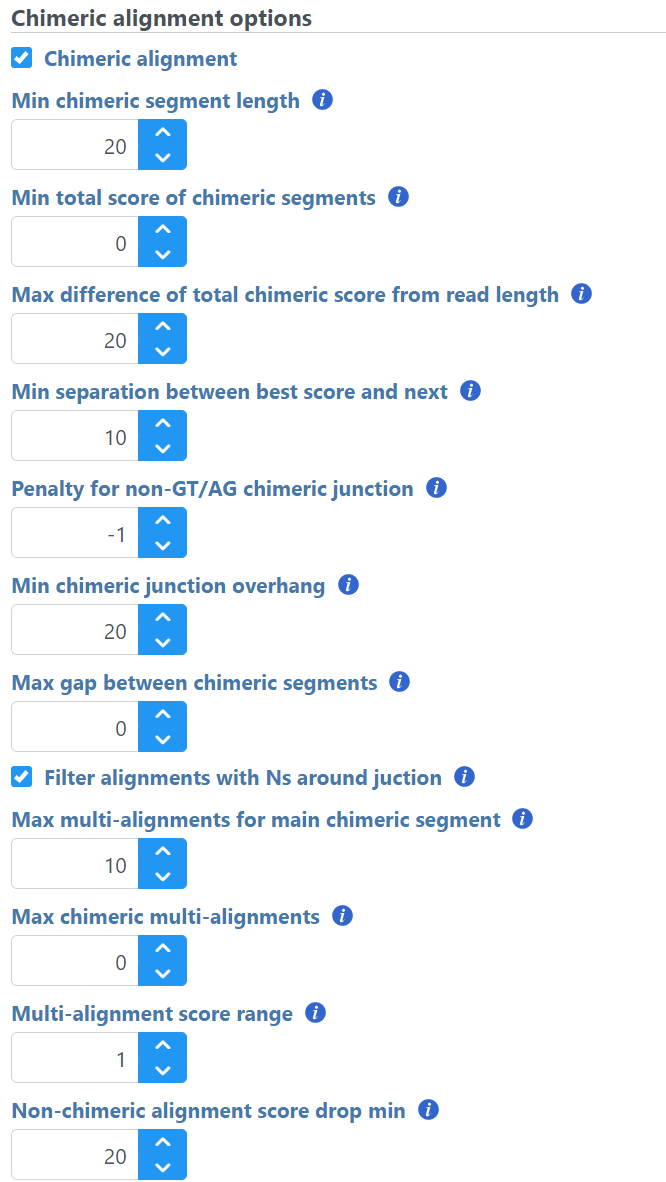
When performing an alignment with STAR, chimeric alignment can be activated by tick-marking the Chimeric alignment option in the Advanced options of the aligner (the Advanced options dialog is reached via the Configure link in the setup dialog). When the Chimeric alignment checkbox is selected, additional options specific to the fusion search algorithm are shown (Figure 11). For a discussion on the details of the options, see STAR documentation.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
First, the genome build that should be used for fusion gene detection needs to be specified (Figure 2).
| |
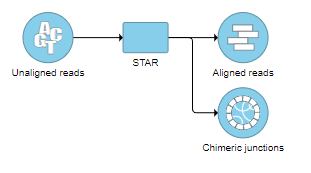
The output is associated with the Chimeric junctions data node (Figure 12), which is a part of the STAR results in addition to Aligned reads node and, optionally, Unaligned reads node.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The next dialog (Fusion options; Figure 3) allows for optimization of several parameters. Min distance between ends specifies the minimum distance (bp) between first in pair and second in pair reads to be considered for a fusion event, while Window gap (bp) defines the minimum distance that needs to be detected between two neighboring fusion candidates in order to label them as independent fusion events. The Annotation model is required to annotate the components of the fusion gene in the output table (see below).
| |||
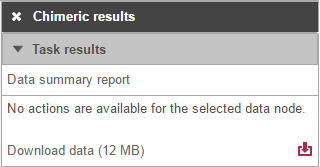
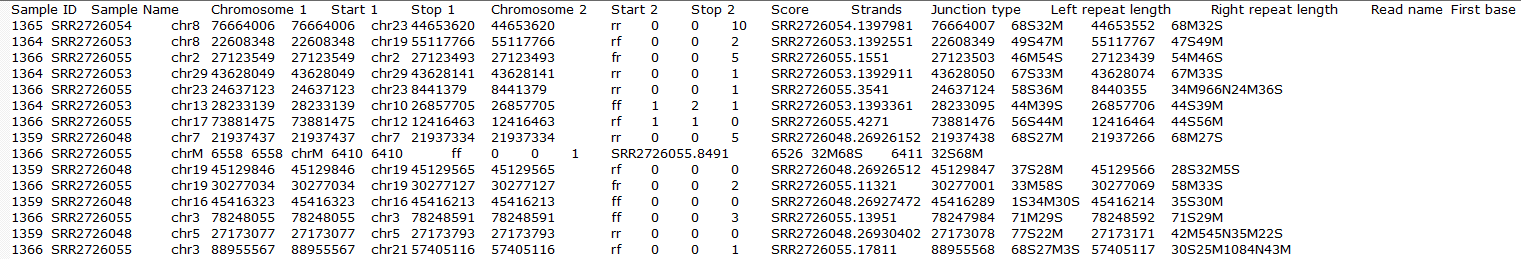
To obtain a .fusion file that summarizes the chimeric reads across samples, select the Chimeric results data node and click Download data in the toolbox (Figure 13). The file is human-readable and can be opened in a text editor (example in Figure 14). For details refer to STAR's documentation.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
As a result, a new data node (Fusion) will be created (Figure 4). Selecting the Fusion node opens the toolbox and the list of fusion genes can then be reached via the Task report link.
| |
Running STAR-Fusion on Chimeric results
STAR-Fusion v1.10 is wrapped into Partek Flow. STAR-Fusion will process the chimeric output generated by the STAR aligner to map junction reads and spanning reads to a reference annotation set. To run fusion detection, select the Chimeric results data node and choose STAR-Fusion from the Variant analysis menu in the toolbox (Figure 15).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
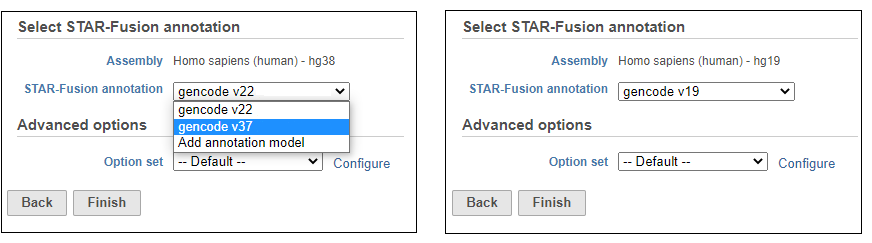
Choose the STAR-Fusion annotation from the drop-down list. We provide automatic downloads of the plug-n-play libraries distributed by Trinity Cancer Transcriptome Analysis Toolkit (CTAT) for Human hg38 (Gencode v22 and v37) and hg19 (Gencode v19) assemblies (Figure 16). If you wish to add your own STAR-Fusion library, you can either import a pre-build CTAT library or gather the appropriate files and build it in Partek Flow. See here for more details on the files you need.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
| |
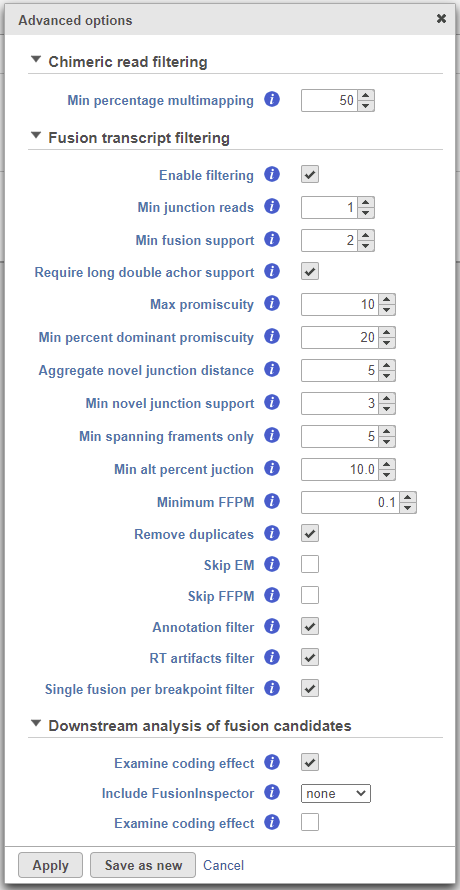
To change any of the advanced options, click the Configure link (Figure 17). To run the task, click Finish.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
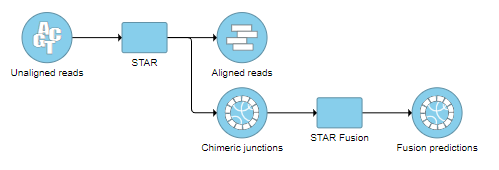
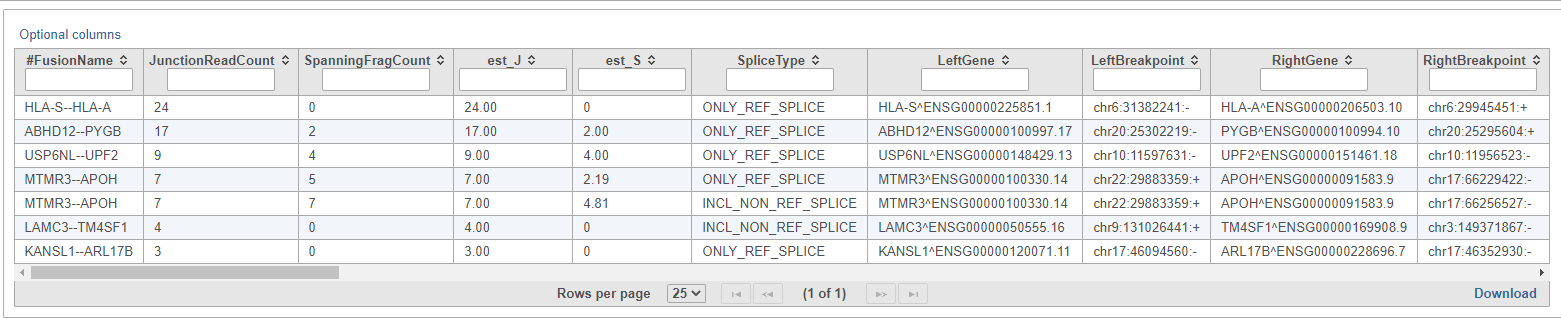
The resulting Fusion predictions task node (Figure 18) can be downloaded to your local machine by selecting the data node and clicking Download data from the toolbox. There will be one tab-separated (.tsv) file per sample. To view the full table, double-click the new data node to open the task report (Figure 19). Each row of the table is a
...
fusion event
...
and the columns
...
contain information about each detected fusion.
...
- Chromosome1FusionName: chromosome ID for the gene on the left side of the fusion;
- Start1: start position of the segment on the left;
- Stop1: stop position of the segment of the left;
- Chromosome 2: chromosome ID for the gene on the right side of the fusion;
- Start2: start position of the segment on the right;
- Stop2: stop position of the segment on the left;
- Sample ID: sample in which the fusion event was identified;
- Counts: number of supporting reads;
- p-value: p-value for the chi-squared test comparing the observed number of counts against the expected number (background distribution);
- Gene1: gene on the left side of the fusion;
- Gene2: gene on the right side of the fusion.
All the columns can be sorted by using the arrow buttons () in column headers.
- the name of the fusion event, given as LeftGene--RightGene. Multiple fusion events can be detected across the same pair of genes, so the FusionName of an event is not necessarily unique;
- JunctionReadCount: indicates the number of RNA-Seq fragments containing a read that aligns as a split read at the site of the putative fusion junction;
- SpanningFragCount: indicates the number of RNA-Seq fragments that encompass the fusion junction such that one read of the pair aligns to a different gene than the other paired-end read of that fragment;
- est_J: estimated junction read counts corrected for multiple mappings;
- est_S: estimated spanning fragment counts corrected for multiple mappings;
- SpliceType: indicates whether the proposed breakpoint occurs at reference exon junctions as provided by the reference transcript structure annotations (Gencode);
- LeftGene: name of the first (left) gene;
- LeftBreakpoint: genome coordinates for the breakpoint in left gene;
- RightGene: name of the second (right) gene;
- RightBreakpoint: genome coordinates for the breakpoint in right gene;
- JunctionReads: sequence identifiers for all junction reads;
- SpanningFrags: sequence identifiers for all spanning fragments;
- LargeAnchorSupport: indicates whether there are split reads that provide 'long' (set to 25bp) alignments on both sides of the putative breakpoint;
- FFPM: fusion fragments per million reads
- LeftBreakDinuc: dinucleotide base pairs at the left breakpoint
- LeftBreakEntropy: the Shannon entropy of the 15 exonic bases flanking the left breakpoint
- RightBreakDinuc: dinucleotide base pairs at the right breakpoint
- RightBreakEntropy: the Shannon entropy of the 15 exonic bases flanking the right breakpoint
- annots: provides a simplified annotation for fusion transcript
| Numbered figure captions | ||||||||
|---|---|---|---|---|---|---|---|---|
|
| |||||||
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
TopHat-Fusion Algorithm
General Overview
TopHat-Fusion is a version of TopHat (see Chapter 6.1) with the ability to align reads across fusion points and detect fusion genes resulting from breakage and re-joining of two different chromosomes or from rearrangements within a chromosome (3). It is independent of gene annotation and can discover fusion products from known genes, unannotated splice variants of known genes or completely unknown genes.
The reads are first aligned to the genome and initially, . The unaligned reads resulting from this initial alignment are then split into multiple 25 bp sequences which are, in turn, aligned to the genome by Bowtie. The TopHat-Fusion algorithm then identifies the cases where the first and the last 25 bp segment segments are aligned to either two different chromosomes or two locations on the same chromosome (spacing is defined by the user). The whole read is then used to identify a fusion point. After the initial fusion candidates are defined, all the segments from the initially unaligned reads are realigned against the fusion points (as well as intron boundaries and indels) and the . The resulting alignments are combined to with the full read alignments.
The most up-to-date TopHat-Fusion version implemented in Partek® Flow® when the manual was written (2.1.0.8) focuses on fusions due to chromosomal rearrangements, while fusions resulting from read-through transcription or trans-splicing were not supported. TopHat-Fusion can handle both paired- and single-end reads, but the support of color-space reads is still pending. For details as well as discussion of TopHat-Fusion options, see TopHat-Fusion home page (4).
Running TopHat-Fusion within Partek Flow
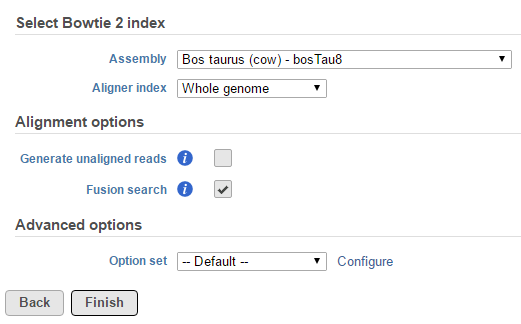
TopHat-Fusion is integrated with in the TopHat 2 task and fusion detection is activated invoked by using the Fusion search check box in the TopHat 2 Alignment options dialog (Figure 61).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
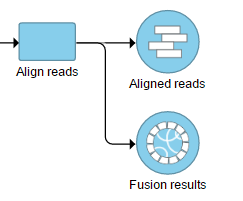
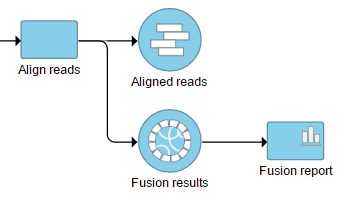
The output is associated with the generated as a new data node Fusion results data node (Figure 7), which is a 2) stemming as part of the if the TopHat 2 results align reads task (in addition to Aligned reads node and, optionally, Unaligned reads node).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
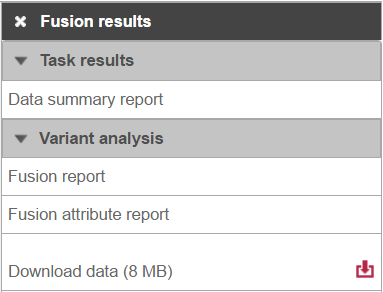
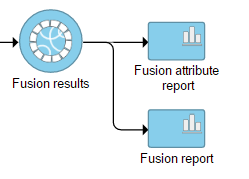
Selecting the Fusion results data node opens the toolboxtask menu, with Variant detection options four options (Figure 8)3): Data summary report, Fusion report, Fusion attribute report, and Download data.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
| |
Clicking the Download data downloads a *.fusion file to the local computer. The file is human-readable and can be opened in a text editor (example in Figure 4). For details refer to TopHat-Fusion documentation.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
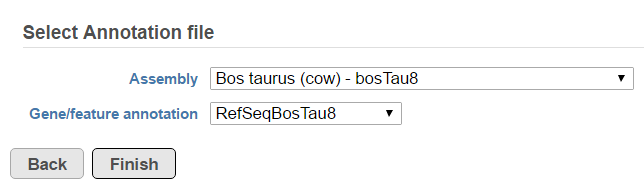
A list of annotated fusion genes, in a form of Fusion report can be obtained by first selecting the Fusion report task node (Figure 2) and then the Task report link from the task menu (Figure 3). Since the task provides an annotated report, an annotation file needs to be specified first (Figure 95).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The result of annotation is the resulting Fusion report task task node as seen in Figure 10.
(Figure 6) can be double-clicked to reveal the full table (Figure 7).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
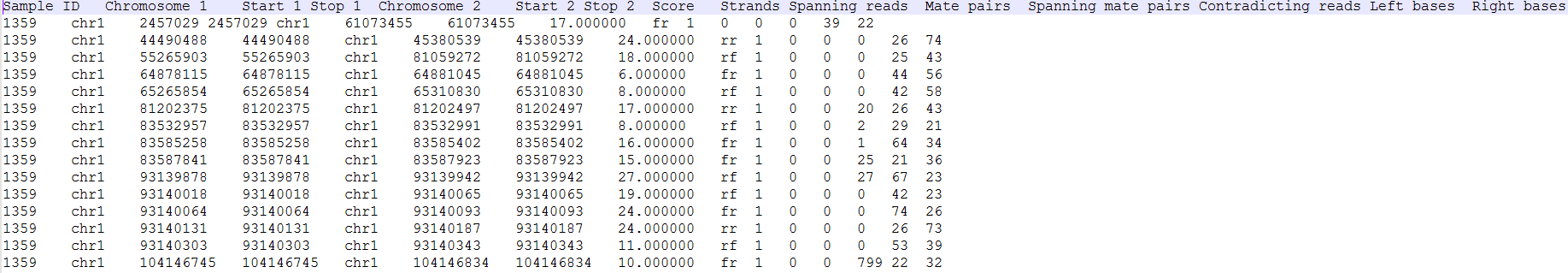
Each row of the table in Figure 11 7 is a potential fusion event, with the columns providing the following information.
- Sample ID: sample in which the fusion event was identified;
- Score: fusion score as defined in the original TopHat-Fusion report (3);
- Type1: genomic section of the left-hand part of the fusion;
- Gene1: gene on the left side of the fusion;
- Transcript1: affected transcript of the Gene1;
- Type2: genomic section of the right-hand part of the fusion;Chromosome 1: chromosome hosting the first (left) segment of the fusion transcript
- Stop 1: end of the first (left) segment of the fusion transcript
- Chromosome 2: chromosome hosting the second (right) part of the fusion transcript
- Start 2: beginning of the second (right) segment of the fusion transcript
- Gene1: gene on the left side of the fusion
- Gene2: gene on the right side of the fusion;Transcript2
- Spanning reads: affected transcript of the Gene2Loci: coordinates of the fusion event (a dash indicates genes on different chromosomes, while a colon indicates that both genes are on the same chromosome with the distance between the parts being given after the colon);
- Strands: orientation of the two chromosomes (e.g. ff indicates that both chromosomes are in forwarding direction);
- Spanning reads: the number of reads spanning the fusion.
- number of reads which were unaligned during the initial phase of TopHat and where only one mate is used as evidence of the fusion event
- Mate Pairs: number of reads which were unaligned during the initial phase of TopHat and where both mates are used as evidence of the fusion event
- Spanning mate pairs: number of reads where both mates were aligned during the initial phase of TopHat, but their pairing is discordant (e.g. different chromosomes, different orientation etc.)
- Contradicting reads: number of reads which do not support the fusion
- Left bases: number of bases on the left side of the fusion
- Right bases: number of bases on the right side of the fusion
All the columns can be sorted by using the arrow buttons () in in column headers, while the type-in boxes can be used for searching.
TopHat-Fusion does not report exact start and stop position for each side of the fusion event. It has a single location for the end of the upstream segment (Stop 1) and the beginning of the downstream segment (Start 2). Therefore, columns Start 1 and Stop 2 are added for (internal) consistency with other Partek Flow tools.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Moreover, Fusion attribute report, when invoked from the Fusion results node, displays a report on attributes of detected fusion genes. Attributes to be tested for association with the fusion should be specified first (Figure 12).
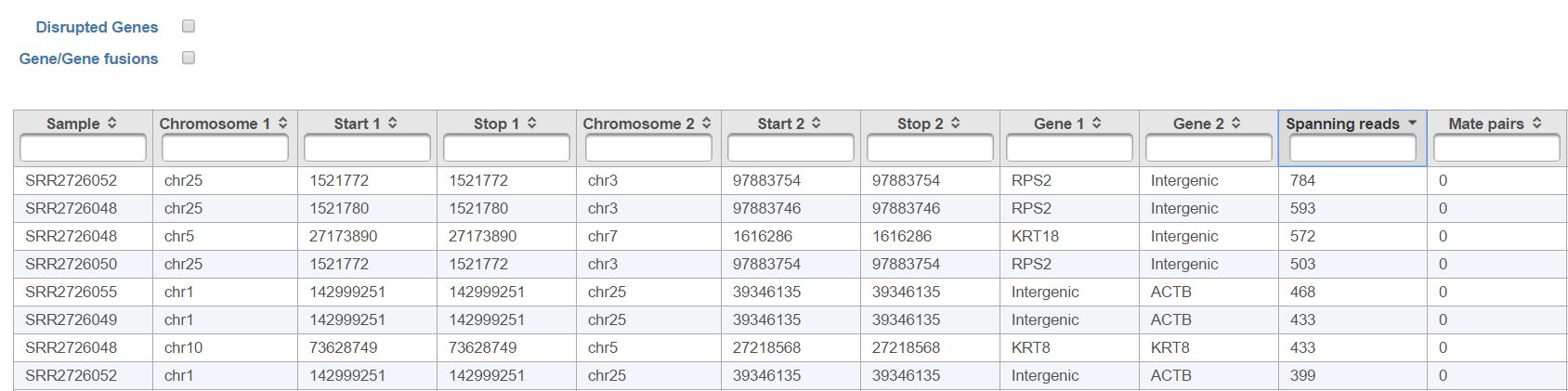
The checkboxes Disrupted Genes and Gene/Gene fusions are filter tools. When selected, Disrupted Genes removes all the rows (fusion events) which have no genes assigned to it, i.e. those that merge two intergenic regions. However, if there is a fusion between a gene and an intergenic region, it will be kept in the table. The Gene/Gene fusions filters in only those fusion events which have an annotated gene on both sides of the breakpoint. In other words, only gene to gene fusions are kept in the table.
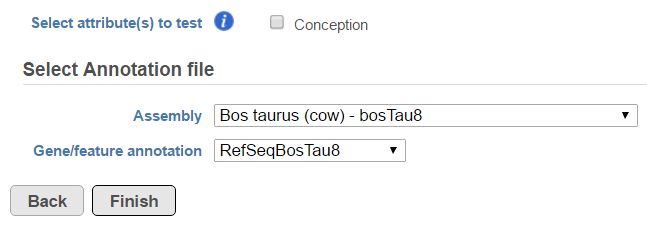
Another table which can be generated based on a Fusion results node is the Fusion attribute report (Figure 3). When the option is selected, it brings up the dialog shown in Figure 8. First, you need to specify one or more categorical attributes (Select attribute(s) to test), which have at least two categories (see Data tab). Second, you need to specify an annotation file, using the Assembly and Gene/feature annotation drop-down lists.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
A new data node, Fusion attribute report, is generated in the Analysis tab (Figure 139) and it provides access to the Task report link in the toolboxtask menu.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
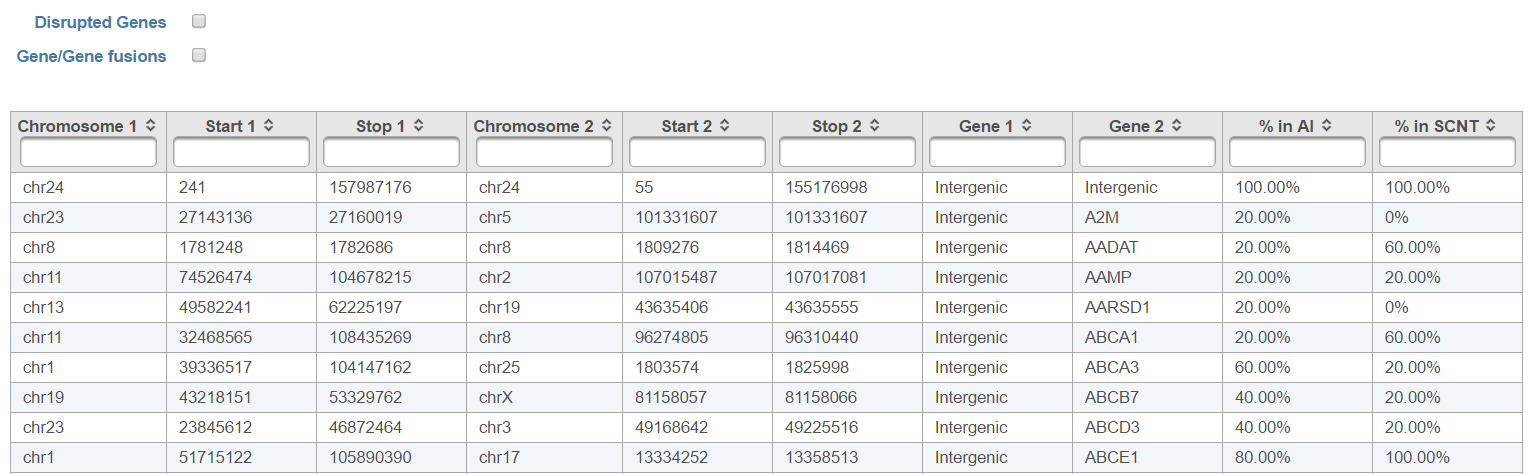
The output, Fusion report table (Figure 1410) resembles the basic TopHat-Fusion output (Figure 117); each row of the table is a single fusion event and three right-most columns are as follows:
- p-value: p-value for the chi-squared test comparing the observed number of counts against across the levels of the attribute specified in the setup;
- % in (attribute level): fraction of reads detected within the samples belonging to the specified level of the attribute (each level is presented as a single column).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
STAR Algorithm
General Overview
STAR aligner (see Chapter 6.1) also has the ability to detect fusion genes (referred to as “chimeric alignments”) (5). During the first phase of alignment, STAR searches for maximal mappable prefixes (seeds) of sequencing reads. In the second phase, all the seeds that align within user-defined genomic windows are stitched together. If an alignment within one genomic window does not cover the entire read sequence, STAR will try to find two or more windows that cover the entire read. This essentially results in detection of fusion events, with different parts of reads aligning to distal genomic locations, or different chromosomes, or different strands.
The most up to date STAR version implemented in Partek Flow when the manual was written (2.3.0) aligns both paired- and single-end reads. Color-space reads are not supported.
Running STAR Chimeric Alignment within Partek Flow
STAR fusion detection algorithm is integrated with STAR aligner and fusion detection is activated by tick-marking Chimeric alignment option in the Advanced options of the aligner (the Advanced options dialog is reached via Configure link in the setup dialog). As soon as the Chimeric alignment is selected, additional options, specific to the fusion search algorithm, are shown (Figure 15). For discussion on the options details, see STAR documentation.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The output is associated with the Chimeric results data node (Figure 16), which is a part of STAR results (in addition to Aligned reads node and, optionally, Unaligned reads node).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Selecting the Chimeric results node opens the toolbox, with Variant detection options (Figure 17).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Fusion report displays an annotated report on detected fusion genes. For that purpose an annotation file needs to be specified first (Figure 18).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The result of annotation is the Fusion report task node as seen in Figure 19.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The list of annotated fusion genes, in a form of Fusion report (Figure 20), can be obtained by first selecting the Fusion report task node and then the Task report link from the toolbox. Each row of the table in Figure 20 is a potential fusion event, with the columns providing the following information.
...
event while the information on the merged segments is on the columns.
- Chromosome 1: chromosome hosting the first (left) segment of the fusion transcript;
- Start 1: beginning of the first (left) segment of the fusion transcript;
- Stop 1: end of the first (right) segment of the fusion transcript;
- Chromosome 2: chromosome hosting the second (right) segment of the fusion transcript;
- Start 2: beginning of the second (right) segment of the fusion transcript;
- Stop 2: end of the second (left) segment of the fusion transcript;
- Gene1: gene on the left side of the fusion;
- Transcript1: affected transcript of the Gene1;
- Type2: genomic section of the right-hand part of the fusion;
- Gene2: gene on the right side of the fusion;
- Transcript2: affected transcript of the Gene2 Loci: coordinates of the fusion event (a dash indicates genes on different chromosomes, while a colon indicates that both genes are on the same chromosome with the distance between the parts being given after the colon);
- Strands: orientation of the two chromosomes (e.g. ff indicates that both chromosomes are in forwarding direction);
- Spanning reads: the number of reads spanning the fusion.
All the columns can be sorted by using the arrow buttons () in column headers.
Figure 20: Fusion report of STAR’s chimeric alignment fusion gene detection algorithm. Each row represents a fusion gene candidate (an example is shown; Score is not applicable to STAR)
Moreover, Fusion attribute report, when invoked from the Chimeric results node, displays a report on attributes of detected fusion genes. Attributes to be tested for association with the fusion should be specified (Figure 21).
Figure 21: Selecting attributes to be tested for association with fusion events (“ER Status” shown as an example)
A new data node, Fusion attribute report, is generated in the Analysis tab (Figure 22) and it provides access to the Task report link in the toolbox.
Figure 22: Fusion attribute report node as a result of annotating Chimeric results generated by STAR’s chimeric alignment algorithm
The output, Fusion report table (Figure 23) resembles the basic TopHat-Fusion output (Figure 11); each row of the table is a single fusion events and three right-most columns are as follows:
- p-value: p-value for the chi-squared test comparing the observed number of counts against across the levels of the attribute specified in the setup;
- % in (attribute level): fraction of reads detected within the samples belonging to the specified level of the attribute (each level is presented as a single column).
...
- % in (category name): fraction of samples within the category with the fusion event.
The checkboxes Disrupted Genes and Gene/Gene fusions are filter tools. When selected, Disrupted Genes removes all the rows (fusion events) which have no genes assigned to it, i.e. those that merge two intergenic regions. However, if there is a fusion between a gene and an intergenic region, it will be kept in the table. The Gene/Gene fusions filters in only those fusion events which have an annotated gene on both sides of the breakpoint. In the other words, only gene to gene fusions are kept in the table.
| Numbered figure captions | ||
|---|---|---|
|
...
|
...
|
...
...
| |||
References
- Annala MJ, Parker BC, Zhang W, Nykter M. Fusion genes and their discovery using high throughput sequencing. Cancer Lett. 2013;340:192-200.
- Costa V, Aprile M, Esposito R, Ciccodicola A. RNA-Seq and human complex diseases: recent accomplishments and future perspectives. Eur J Hum Genet. 2013;21:134-142.
- Kim D, Salzberg SL. TopHat-Fusion: an algorithm for discovery of novel fusion transcripts. Genome Biology. 2011;12:R72
- TopHat-Fusion. An algorithm for discovery of novel fusion transcripts. http:// http://tophat.cbcb.umd.edu/fusion_index.html Accessed on April 25, 2014
- Dobin A, Davies CA, Schlesinger F et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15-21.
| Additional assistance |
|---|
|
| Page Turner |
|---|
- Haas B.J, Dobin A, Li B. et al. Accuracy assessment of fusion transcript detection via read-mapping and de novo fusion transcript assembly-based methods. Genome Biol. 2019;20:213 (2019)
| Additional assistance |
|---|
| Rate Macro | ||
|---|---|---|
|
...