Page History
...
Multi-omics single cell analysis is based on simultaneous detection of different types of biological molecules on the same cells. Common multi-omics techniques include feature barcoding or CITE-seq (cellular indexing of transcriptomes and epitopes by sequencing) technologies, which enable parallel assessment of gene and protein expression. Specific bioinformatics tools have been developed to enable scientists to integrate results of multiple assays and learn relative importance of each type (or each biological molecule) in identification of cell types. Partek Flow supports weighted nearest neighbour (WNN) analysis (1), which can help combine output of two or more molecular assays.
Invoking Find Multimodal Neighbours
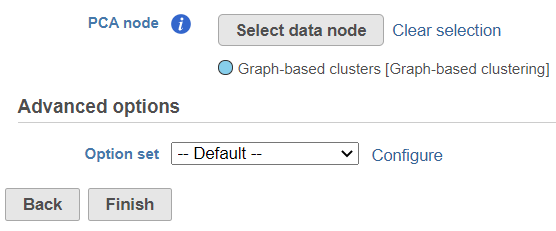
To start, select a PCA data node of one of the assays (e.g. gene expression) and go to Exploratory analysis > Find multimodal neighbours in the toolbox. On the task setup page, use the Select data node button to point to the PCA data node of the other assay (e.g. protein expression) (Figure 1).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
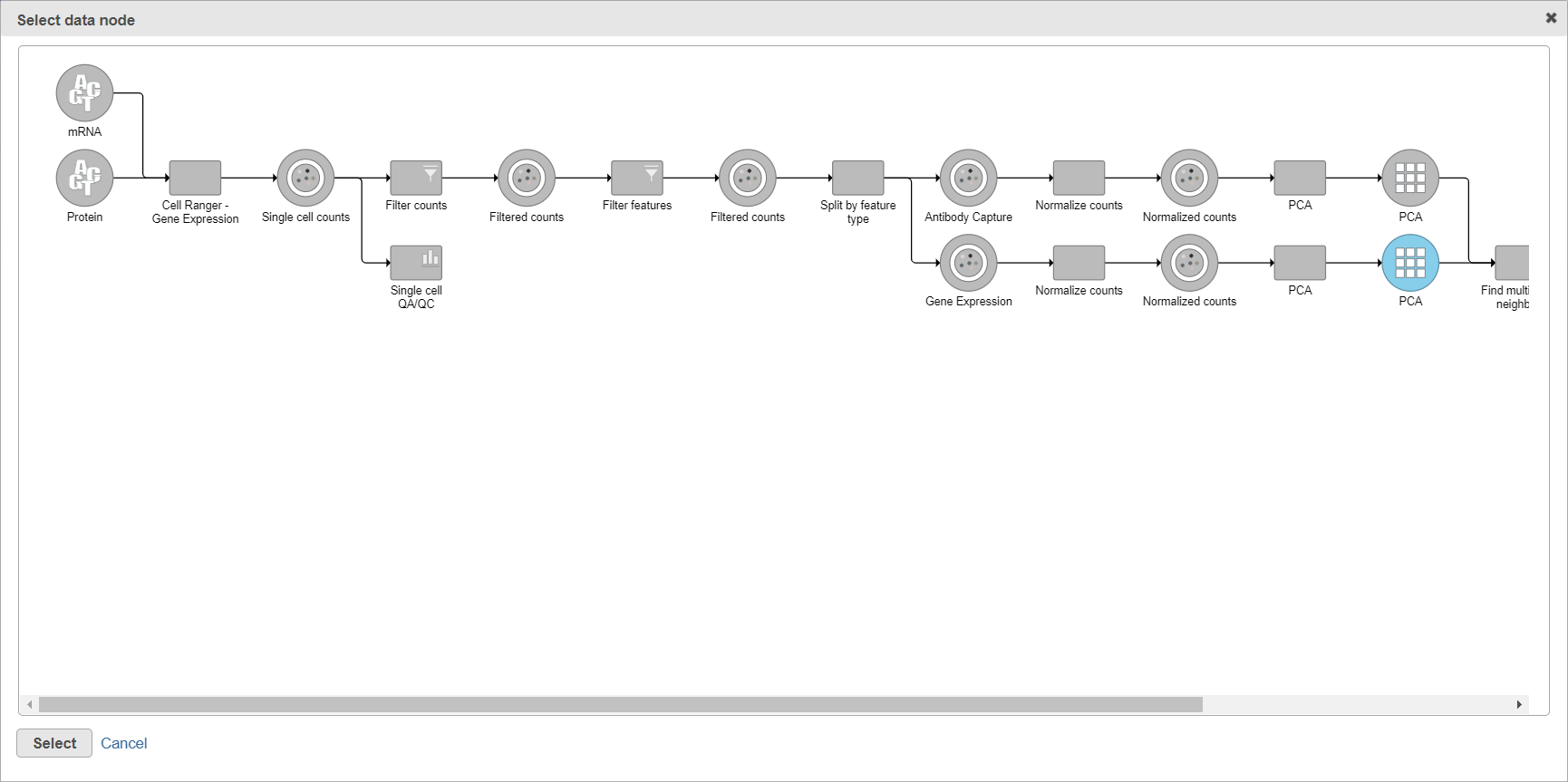
When you click the Select data node button, Partek Flow will open another dialog, showing your current pipeline (Figure 2). Data nodes that can be used for WNN are in blue. To pick a node, left-click on it and then push the Select button.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
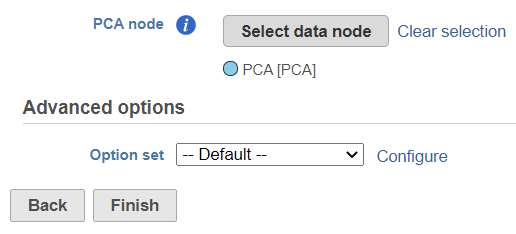
The selected data node is shown under the Select data node button. If you made a mistake, use the Clear selection link (Figure 3).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
References
- Hao Y, Hao S, Andersen-Nissen E, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184(13):3573-3587.e29. doi:10.1016/j.cell.2021.04.048
...