Page History
...
This user guide illustrates:
| Children List |
|---|
Section Heading
Section headings should use level 2 heading, while the content of the section should use paragraph (which is the default). You can choose the style in the first dropdown in toolbar.
This user guide assumes the user is familiar with the hierarchy of spreadsheets and analysis in Partek® Genomics Suite®.
Many plots available in Partek Genomics Suite are not discussed in this user guide. A more thorough review of Partek Genomics Suite visualizations can be found in Chapter 6: The Pattern Visualization System of the Partek User's Manual available from Help > User’s Manual in the Partek Genomics Suite main toolbar.
There is no specific data set for this tutorial; you may use one of your own microarray experiments or use a data set from one of our tutorials.
Visualizations are generated using data from a spreadsheet. Some visualizations allow interactive filtering on the plot, but others do not. If you only wish to include certain rows or columns in a visualization, you may need to create a spreadsheet with only the rows or columns of interest by applying a filter and cloning the spreadsheet.
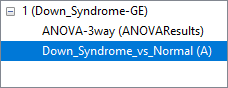
In general, gene intensity values may be visualized from either an ANOVA spreadsheet or a filtered ANOVA spreadsheet. Because probe intensity data is stored in the parent spreadsheet, the parent and child spreadsheets should be visible in the spreadsheet navigator with the appropriate parent/child relationship (Figure 1).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|