Page History
...
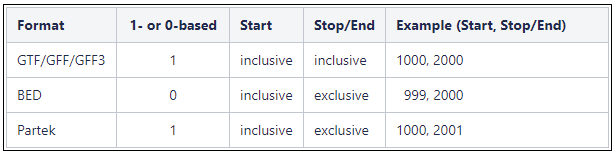
Genome coordinates for annotation models stored in Partek Flow are 1-based, start-inclusive, and stop-exclusive. This means that the first base position starts from one, the start coordinate for a feature is included in the feature and the stop/end coordinate is not included in the feature. These are the genome coordinates that are printed in various task reports and output files when an annotation model is involved in the task. When custom annotation files are added to Partek Flow, the genome coordinates are converted into this format. The coordinates are converted back if necessary for a specific task. shows how the genome coordinates vary between different annotation formats.
Can I add transgenes to my reference files?
...
Fold change indicates the extent of increase or decrease in feature expression in a comparison. In Partek Flow, fold change is in linear scale (even if the input data is in log scale). It is converted from ratio, which is the LSmean of group one divided by LSmean of group two in your comparison. When the ratio is greater than 1, fold change is identical to ratio; when the ratio is less than 1, fold change is -1/ratio. There is no fold change value between -1 to 1. When ratio/fold change is 1, that means there is no change between the two groups.
Log ratio option in Partek Flow is converted from ratio, this is a value comparable to log fold change in some other tools.
Can I label a Volcano plot with gene names?
...