...
| Numbered figure captions |
|---|
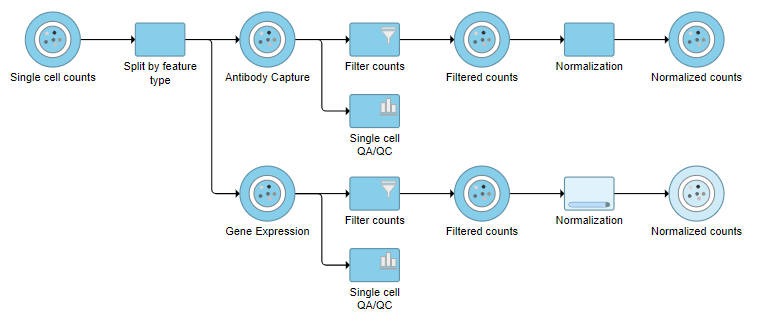
| SubtitleText | The two normalization tasks produce Normalized counts data nodes |
|---|
| AnchorName | Normalized counts output |
|---|
|
|
Merge Protein and mRNA data
For quality filtering and normalization, we needed to have the two data types separate as the processing steps were distinct. For downstream analysis, we want to be able to analyze protein and mRNA data together. To bring the two data types back together, we will merge the two normalized counts data nodes.
- Click the Normalized counts data node on the Antibody Capture branch of the pipeline
- Click Pre-analysis tools in the toolbox
- Click Merge matrices
- Click Select data node to launch the data node selector
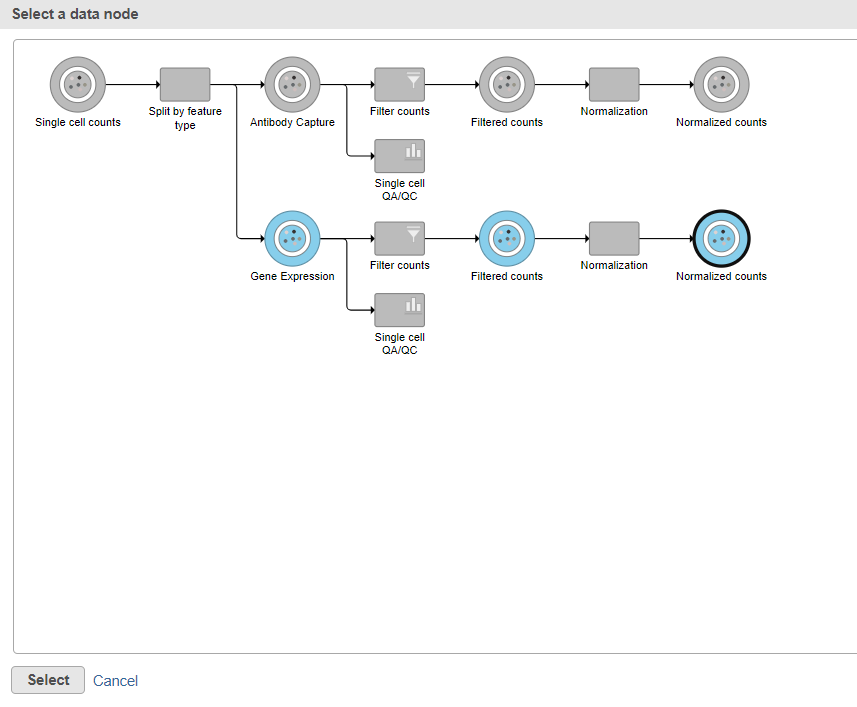
Data nodes that can be merged with the Antibody Capture branch Normalized counts data node are shown in color (Figure ?).
| Numbered figure captions |
|---|
| SubtitleText | Select the normalizated gene expression counts to merge the protein counts with |
|---|
| AnchorName | Merge protein and gene expression counts |
|---|
|
|
- Click the Normalized counts data node on the Gene Expression branch of the pipeline
- Click Select
- Click Finish to run the task
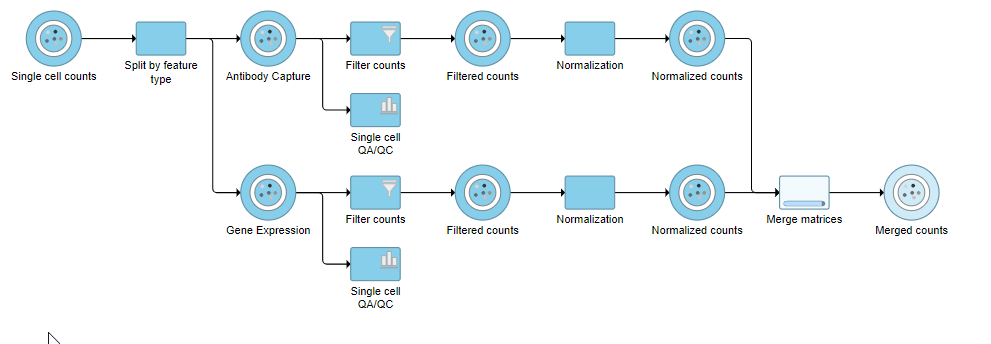
The output is a Merged counts data node (Figure ?). This data node will include the normalized counts of our protein and mRNA data. The intersection of cells from the two input data nodes is retained so only cells that passed the quality filter for both protein and mRNA data will be included in the Merged counts data node.
| Numbered figure captions |
|---|
| SubtitleText | Merged counts output |
|---|
| AnchorName | Merged counts output |
|---|
|
|
Collapsing tasks to simplify the pipeline
To simplify the appearance of the pipeline, we can group task nodes into a single collapsed task. Here, we will collapse the filtering and normalization steps.
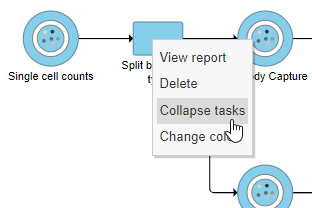
- Right-click the Split by feature type task node
- Choose Collapse tasks from the pop-up dialog (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Choosing the first task node to generate a collapsed task |
|---|
| AnchorName | Collapse tasks |
|---|
|
|
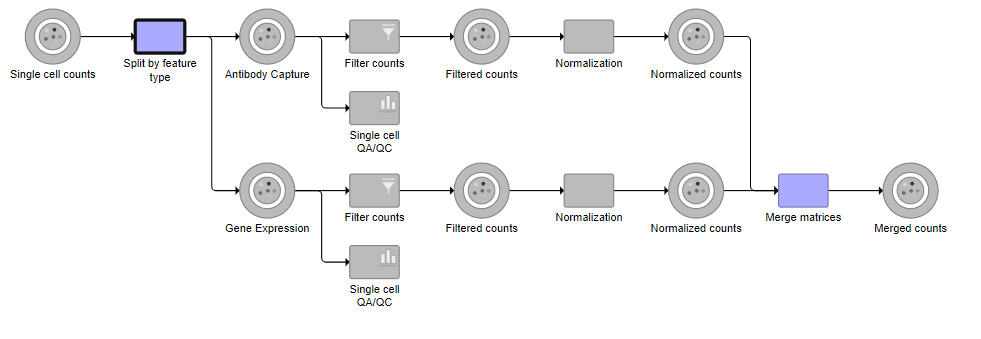
Tasks that can be selected for the beginning and end of the collapsed section of the pipeline are highlighted in purple (Figure ?). We have chosen the the Split matrix task as the start and we can choose Merge matrices as the end of the collapsed section.
| Numbered figure captions |
|---|
| SubtitleText | Tasks that can be the start or end of a collapsed task are shown in purple |
|---|
| AnchorName | Available tasks to collapse |
|---|
|
|
- Click the Merge matrices task to choose it as the end of the collapsed section
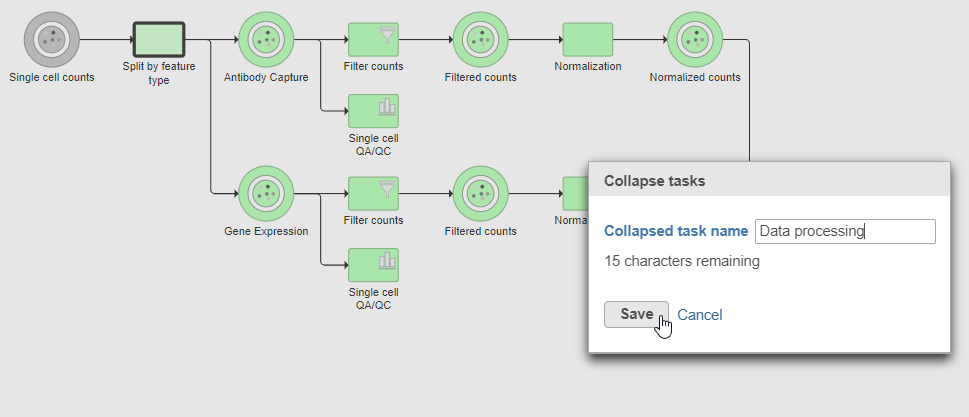
- Name the Collapsed task Data processing
- Click Save (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Naming the collapsed task |
|---|
| AnchorName | Save collapsed task |
|---|
|
|
...
| Numbered figure captions |
|---|
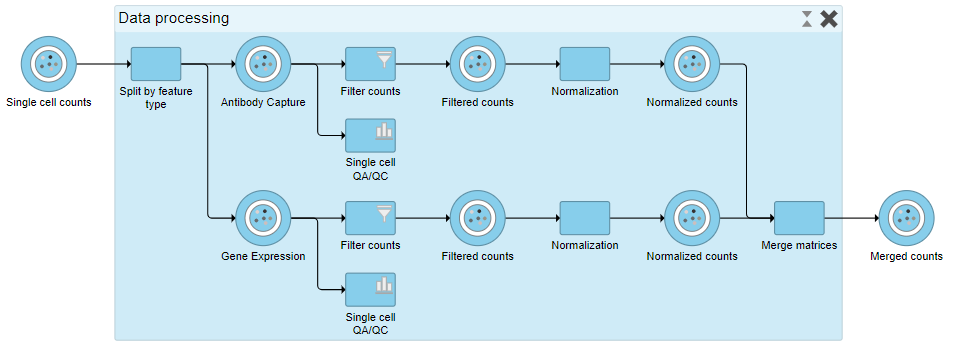
| SubtitleText | Expanding a collapsed task to show its components |
|---|
| AnchorName | Expanding collapsed task |
|---|
|
|
To re-collapse the task, you can double click the title bar or click the  Image Modified icon in the title bar. To remove the collapsed task, you can click the
Image Modified icon in the title bar. To remove the collapsed task, you can click the  Image Modified. Please note that this will not remove tasks, just the grouping.
Image Modified. Please note that this will not remove tasks, just the grouping.
- Double-click the Data processing title bar to re-collapse
References
[1] Stoeckius, M., Hafemeister, C., Stephenson, W., Houck-Loomis, B., Chattopadhyay, P. K., Swerdlow, H., ... & Smibert, P. (2017). Simultaneous epitope and transcriptome measurement in single cells. Nature methods, 14(9), 865.
...






