Page History
...
Cells undergo changes to transition from one state to another as part of development, disease, and throughout life. Because these changes can be gradual with transitional stages between endpoints, trajectory analysis attempts to describe the progress through a biological process as a position along a path. Because biological processes are rarely as simple as a direct line from one starting point and one end pointoften complex, trajectory analysis builds branching trajectories where different paths can be chosen at different points along the trajectory. Progress along the The progress of a cell along a trajectory from the root or starting point, is described by the starting point or root, can be quantified as a numeric value, pseudotime.
In Partek Flow, we use tools from Monocle 2 (Qui et al. 2017) to build trajectories, identify states and branch points, and calculate pseudotime values. The output of Trajectory analysis includes an interactive scatter plot visualization for viewing the trajectory and setting the root state (starting point of the trajectory) and adds a categorical cell level attribute, State. From the Trajectory analysis task report, you can run a second task, Calculate pseudotime, which adds a numeric cell-level attribute, Pseudotime, which is calculated using the chosen root state. Using the state and pseudotime attributes, you can perform downstream analysis to identify genes that change over pseudotime and characterize branch points.
Prerequisites for trajectory analysis
For Trajectory analysis to work as expected, there are a few things you do prior to running it.Prior to performing trajectory analysis, you should:
1) Normalize the data
Trajectory analysis requires normalized counts as the input data. We recommend our default "CPM, Add 1, Log 2" normalization for most scRNA-Seq data.
...
The trajectory should be built using a set of genes that increase or decrease as a function of progression through the biological processes being modeled. One example is using differentially expressed genes between cells collected at the beginning of the process to cells collected at the end of the process. If you have no prior knowledge about the process being studied, you can try identifying genes that are differentially expressed between clusters of cells or genes that are highly variable within the data set. Generally, you should try to filter to 1,000 to 3,000 informative genes prior to running Trajectory analysis. The list manager functionality in Partek Flow is useful for creating a list of genes to use in the filter.
Parameters
Two parameters are exposed for trajectory analysis. First, the number of dimensions. Dimensionality of the reduced space
While the trajectory is always visualized in a 2D scatter plot, the underlying structure of the trajectory may be more complex and better represented by more than 2 dimensions. Second, you
Scaling
You can choose to scale the genes prior to building the trajectory. Scaling removes any differences in variability between genes, while not scaling allows more variable genes to have a greater weight in building the trajectory.
Task report
The Trajectory analysis task report is a 2D scatter plot (Figure 1).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The trajectory is shown with a black line showing the trajectory, numbers at each branch point, and the cells . Branch points are indicated by numbers in black circles. By default, cells are colored by state. Like any scatter plot, you can color
You can use the control panel on the left to color, size, and shape by genes and attributes to help identify which state is the root or origin of the trajectory (Figure 2). To select a root state, click
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
You can also split by any categorical attribute (Figure 3)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Calculating pseudotime
To calculate pseudotime, you must choose a root state. Click any cell belonging to that state . Clicking Calculate pseudotime will run the task to calculate pseudotime values for each cell using the selected state as the root.
to select the state. The selected state will be bold while unselected cells are dimmed (Figure 4).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
To use the selected state as the Root state for pseudotime calculation, select the state and then click the Calculate pseudotime button (Figure 5).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
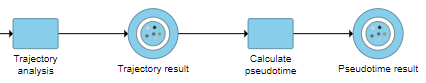
This will run a task on the analysis pipeline, Calculate pseudotime, and output a new Pseudotime result data node (Figure 6).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
References
Xiaojie Qiu, Qi Mao, Ying Tang, Li Wang, Raghav Chawla, Hannah Pliner, and Cole Trapnell. Reversed graph embedding resolves complex single-cell developmental trajectories. Nature methods, 2017.
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|